Research
The research interests of the faculty of the Walter S. and Lucienne Driskill Graduate Program in Life Sciences represents a broad and diverse array of topics in modern biomedical sciences. Learn more about Feinberg scientists' work via the topics below.

Mohamed E Abazeed
Professor, Radiation Oncology
Individualize cancer care (radiotherapy) by helping physicians recommend treatments based on the genetic and imaging features of individual tumors.

Mohamed E Abazeed
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Pharmacology
Precision oncology facilitates individualized treatment decisions on the basis of patient and tumor specific factors for an increasing proportion of cancer patients. Despite growing evidence that inter-patient variation affects treatment responses after radiotherapy, patients receiving these treatments continue to be treated with the same or similar doses. We seek to develop an information capability at the forefront of personalized radiotherapy treatments. We achieve this through the assembly of experimental scaffolds that span the translational research spectrum and help us understand tumor complexity and predict clinical outcomes.
Briefly, we conduct large-scale projects that capture the diversity of our patients and provide a rich substrate for computational and mathematical models of cancer’s propensity to resist our treatments. Three large-scale projects have been completed or are currently in progress including: 1) The X-ray Target Discovery and Development (XTD2) project, which profiled 533 cancer cell line survival comprising 26 cancer types to ionizing radiation. This project represented the largest profiling effort of cancer cell line survival after irradiation ever conducted. 2) The Pan-cancer Radiogenomic Atlas is a gene variant profiling project that interrogated >1000 common and rare genetic variants for response to ionizing radiation in immortalized human cells (non-cancer cells). Current work is building on the unary profiling methodology to study the interaction between varied gene variants, thus building toward greater complexity. 3) The 10,000 Avatar Project was inaugurated by our group in 2019. This will be the largest patient-derived xenograft (PDX) mouse experiment conducted to date by any group. ~10,000 mice engrafted with ~500 genetically annotated PDXs will be irradiated using a singular experimental platform. This work will correlate genetic and other omic (e.g. transcriptomic, metabolomic, et cetera) alterations with the likelihood of response to radiotherapy and matched recurrent tumors.
Concurrent with the large-scale biological profiling approaches described above, we have developed a clinomic dataset that integrates clinical information (e.g. demographics, treatments, outcomes) and patient avatar models (patient-derived xenografts) with omic outputs for individual patients. The latter include radiomics (embedded quantitative data derived from imaging modalities like computed tomography), genomics (genetic information derived from the patient’s tumor or germline), transcriptomics (gene expression), and others. Using this information, we seek to design and implement tools that can augment the physician’s ability to estimate the probability of treatment failures and modulate failure by individualized treatment recommendations.
For lab information and more, see Mohamed Abazeed's, MD,PhD, faculty profile.
Publications
See Dr. Abazeed's publications in PubMed.
Contact Us
Contact the Abazeed Lab at 312-503-2195. You may also contact Dr. Abazeed directly via email.

Mohamed Abdel Mohsen
Associate Professor, Medicine (Infectious Diseases)
Glycoimmunology and Disease Pathogenesis

Mohamed Abdel Mohsen
Research Area(s): Immunology; Microbiology
My laboratory investigates how alterations in cellular glycosylation regulate immune responses in the contexts of HIV infection, cancer, and aging. We integrate glycomics, immunology, virology, and systems biology to uncover how glycan-mediated communication shapes inflammation, immune dysfunction, and disease progression. Our goal is to translate these insights into new therapeutic strategies that restore immune balance and promote healthy aging.
For lab information and more, see Mohamed Abdel-Mohsen's faculty profile or the Abdel-Mohsen lab website.
Publications
See Dr. Abdel-Mohsen's publications in PubMed.
Contact Us
Contact Dr. Abdel-Mohsen.

Sarki A Abdulkadir
Professor, Urology, Pathology
Mechanisms of prostate cancer initiation, progression and recurrence and strategies to therapeutically target these processes

Sarki A Abdulkadir
Research Area(s): Cancer Biology; Developmental and Stem Cell Biology; Pharmacology; Reproductive Biology
Our laboratory focuses on understanding the molecular mechanisms that drive prostate cancer initiation, progression and recurrence with the ultimate goal of developing therapeutic strategies that target these processes. Our approach includes the genomic analysis of human tumors, cell culture studies and the use of genetically engineered mouse models. We have a strong interest in genomics and gene regulation, oncogenic kinases as potential molecular therapeutic targets and the use of in vivo lineage tracing to define the fates of specific cell populations in tumorigenesis.
Specific projects include:
The role of the oncogenic serine/threonine kinase PIM1 in prostate cancer - PIM1 is coexpressed with c-MYC and dramatically enhances c-MYC-driven prostate tumorigenesis in a kinase-dependent manner. Notably, PIM1 is induced in tumors by hypoxia, radiation and treatment with docetaxel, a common but largely ineffective option for patients with advanced castration-resistant prostate cancer. PIM1 induction by hypoxia/radiation/docetaxel promotes prostate cancer cell survival and therapeutic resistance. Therefore, PIM1 may represent a valuable therapeutic target in prostate cancer. We are using new mouse models of prostate cancer for testing the efficacy of novel PIM1 kinase inhibitors in treating prostate cancer and reversing therapeutic resistance. We have also identified novel candidate PIM1-interacting proteins in prostate epithelial cells. Among the proteins identified are a MYC transcriptional cofactor and a prostate stem cell marker/regulator. We are investigating how PIM1 promotes prostate tumorigenesis by phosphorylating these substrates involved in regulating MYC transcriptional activity and stem cell function.
Cellular and molecular determinants of prostate cancer recurrence - A major clinical problem in prostate cancer is that of tumor recurrence following initial apparently successful therapy. Recurrent tumors may arise from a small number of "cancer stem-like cells" that survive the initial therapeutic intervention and have the capacity to regenerate the tumor. We are using lineage tracing to examine the competence of specific prostate epithelial cell types to regenerate tumors following therapy in mice.
Targeting lethal prostate cancer – We are using our mouse model of lethal prostate cancer based on alterations in Myc, Pten and Tp53 to develop new targeted therapies. One current project involves the targeting of EphB4 receptor tyrosine kinase using an antagonist as a therapeutic strategy.
For more information, see Dr. Abdulkadir's faculty profile.
Publications
See Dr. Abdulkadir's full publications in PubMed.
Contact Us
Dr. Abdulkadir
Lab Telephone: 312-503-5031

Mazhar Adli
Professor, Obstetrics and Gynecology (Reproductive Science in Medicine), Cell and Developmental Biology
Preventing cancer development and chemotherapy resistance using genomic and epigenomic approaches

Mazhar Adli
Research Area(s): Cancer Biology; Cell Biology; Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Pharmacology; Reproductive Biology
I am interested in understanding the key drivers of cancer and identifying novel therapeutic drug combinations to prevent cancer development and chemotherapy resistance. To achieve these goals, our lab is using and developing genomic and epigenomic mapping, editing and imaging approach to understand genome regulation in normal and malignant settings. We integrate experimental approaches with large-scale computational data analysis to verify our experimental observations and come up with new testable hypotheses. Our laboratory is utilizing and also developing cutting-edge functional genomics strategies and developing novel CRISPR based manipulation tools to understand dynamic gene regulation and 3D genome organization in normal and malignant settings. These efforts are based on our previous expertise in genome-wide approaches and development of novel technologies for cancer research. Our lab has developed particular expertise in utilizing and developing CRISPR based technologies.
For more information, see Dr. Adli's faculty profile or the Adli lab website.
Publications
See Dr. Adli's publications in PubMed.
Contact Us

Atique U Ahmed
Professor, Neurological Surgery
Implications of cancer cell plasticity on tumor progression and therapy resistance

Atique U Ahmed
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics; Immunology; Neurobiology; Pharmacology
Atique Ahmed, Ph.D., is an Associate Professor in the Department of Neurosurgery at Northwestern University and a distinguished Lurie Comprehensive Cancer Center member. He completed his BSc (Hons) degree at Winona State University in Minnesota and earned his Ph.D. from the Mayo Clinic School of Medicine. He pursued postdoctoral training at the University of Chicago.
Ahmed's lab research focuses on cancer cell plasticity and its implications for tumor progression and therapy resistance. He investigates how cancer cells adapt and alters their characteristics in response to the tumor microenvironment, including their resistance to therapy and potential for metastasis. Dr. Ahmed aims to develop innovative therapies to prevent the transition to more aggressive states and inhibit tumor spread by studying the mechanisms underlying cancer cell plasticity.
One significant aspect of Lab research is exploring epigenetic changes in cancer. Epigenetic alterations can influence gene expression in cell growth and division, rendering cancer cells more aggressive and resistant to treatment. Dr. Ahmed's lab has demonstrated that anti-cancer therapies can induce epigenetic plasticity, enabling cancer cells to adapt and evade treatment. Through his investigations, he seeks to identify strategies that reverse or block these changes, ultimately preventing cancer cells from becoming more aggressive or resistant to therapy.
A recent milestone from Ahmed Lab is the initiation of a clinical trial that investigates the repurposing of Mycophenolate mofetil (MMF), an FDA-approved drug, to target plasticity-driven chemoresistance in brain tumors. The lab's promising findings suggest that MMF can sensitize tumor cells to treatment and potentially improve outcomes for glioblastoma (GBM) patients. By evaluating the safety and efficacy of MMF in a clinical setting, Dr. Ahmed aims to enhance current GBM therapies and pave the way for novel treatment approaches in other cancer types.
For lab information and more, see Atique Ahmed's faculty profile.
Publications
See Dr. Ahmed's publications in PubMed.
Contact Us

Lisa N Akhtar
Assistant Professor, Pediatrics (Infectious Diseases)
Viral virulence in the pediatric brain

Lisa N Akhtar
Research Area(s): Cell Biology; Microbiology; Neurobiology
The Akhtar laboratory strives to understand how viruses infect and cause pathogenesis in the pediatric brain. Our current focus is determining the contribution of viral genetic variability to neurovirulence following neonatal herpes simplex virus (HSV) infection. HSV infection of the neonatal brain causes severe encephalitis and permanent neurologic deficits, but the factors that promote invasive central nervous system (CNS) infection are not known. Recent studies show that substantial genetic variability exists between clinical HSV genomes, but have not evaluated how these variations impact viral growth characteristics or human disease manifestations. We utilize viral whole genome sequencing of clinical HSV isolates to identify viral genetic variations associated with neonatal CNS disease. We then use human and murine neuronal in vitro models to determine their impact on viral growth characteristics, and murine models of encephalitis to determine their ability to alter disease progression. These studies are providing the first insights into how variations in the neonatal HSV genome impact neurovirulence and the development of CNS disease.
For lab information and more, see Dr. Akhtar's faculty profile.
Publications
See Dr. Ahmed's publications in PubMed.
Contact Us

Daniel Arango
Assistant Professor, Pharmacology
Role of post-transcriptional modifications of RNA in the proliferation, differentiation and survival of cancer cells

Daniel Arango
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Genetics, Epigenetics and Genomics; Pharmacology
For more information, see Dr. Arango's faculty profile or the Arango lab website.
Publications
See Dr. Arango’s publications in PubMed.
Contact Us

Rajeshwar Awatramani
Professor, Neurology (Movement Disorders)
Professor, Weinberg College of Arts and Sciences
Dopamine neurogenesis and subtypes and the role of microRNAs in Schwann cell (SC) differentiation.

Rajeshwar Awatramani
Research Area(s): Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology
Topic 1. Mechanisms underlying dopamine neurogenesis
The floor plate, the ventral organizing center in the embryonic neural tube, patterns the neural tube by secreting the potent morphogen Shh. Using genetic fate mapping, we have recently shown that the midbrain floor plate, unlike the hindbrain and spinal cord floor plate, is neurogenic and is the source of midbrain dopamine neurons (Joksimovic, et al, 2009 Nature Neuroscience, Joksimovic et al. 2009 PNAS). We are interested in understanding pathways that are involved in floor plate neurogenesis and production of dopamine neurons. We have shown that Wnt signaling is critical for the establishment of the dopamine progenitor pool and that miRNAs may modulate the dosage and timing of the Wnt pathway (Anderegg et al, PloS Genetics 2013).
Topic 2. Deconstructing Dopaminergic Diversity
The neurotransmitter dopamine, produced mainly by midbrain dopaminergic neurons, influences a spectrum of behaviors including motor, learning, reward, motivation and cognition. In accordance with its diverse functions, dopaminergic dysfunction is implicated in a range of disorders affecting millions of people, including Parkinson’s disease (PD), schizophrenia, addiction and depression. How a small group of neurons underpins a gamut of key behaviors and diseases remains enigmatic. We postulated that there must exist several molecularly distinct dopaminergic neuron populations that, in part, can account for the plethora of dopaminergic functions and disorders. We are currently working to test this hypothesis and define dopamine neuron subtypes.
Topic 3. MicroRNAs in Schwann cell (SC) differentiation
MicroRNAs, by modulating gene expression, have been implicated as regulators of various cellular and physiological processes including differentiation, proliferation and cancer. We have studied the role of microRNAs in Schwann cell (SC) differentiation by conditional removal of the microRNA processing enzyme, Dicer1 (Yun et al, 2010, J Neurosci) . We reveal that mice lacking Dicer1 in SC (Dicer1 cKO) display a severe neurological phenotype resembling congenital hypomyelination. SC lacking Dicer1 are stalled in differentiation at the promyelinating state and fail to myelinate axons. We are beginning to determine the molecular basis of this phenotype. Understanding this will be important not only for congenital hypomyelination, but also for peripheral nerve regeneration and SC cancers.
For more information, please see Dr. Awatramani's faculty profile.
Publications
View Dr. Awatramani's complete list of publications in PubMed.
Contact Us
Dr. Awatramani, PhD at 312-503-0690

Elnur Babayev
Assistant Professor, Obstetrics and Gynecology (Reproductive Endo and Infertility)
Modulation of factors affecting mammalian reproduction

Elnur Babayev
Research Area(s): Cell Biology; Genetics, Epigenetics and Genomics; Reproductive Biology
The Babayev lab is interested in understanding the determinants of gamete quality and the mechanisms of reproductive aging process from birth to menopause, investigating the environmental factors that affect gamete quality and reproductive longevity, and in bringing new therapeutic modalities from bench to bedside. Our current research focuses include: 1) the investigation of the egg quality of adolescent patients undergoing fertility preservation via the analysis of the oocyte microenvironment 2) the identification of environmental factors that impact reproduction and understanding their mechanisms of action. This include the impact of endocrine disrupting chemicals on reproductive aging and the effects of the changing environment on reproductive potential. 3) testing the feasibility of gene therapy in human reproductive tissues to treat reproductive system diseases that currently are thought to be incurable.
For more information, see Dr. Bababyev's faculty profile.
Publications
View Dr. Babayev's complete list of publications in PubMed.
Contact Us

Kelly E Bachta
Assistant Professor, Medicine (Infectious Diseases)
Antimicrobial resistance mechanisms and pathogenesis of clinically-important bacterial pathogens including Pseudomonas aeruginosa and Enterococcus faecium

Kelly E Bachta
Research Area(s): Biochemistry and Structural Biology; Lung Biology; Microbiology; Neurobiology
The Bachta laboratory has two main research foci:
- We investigate the pathogenesis of Pseudomonas aeruginosa infections using imaging and sequencing techniques to define infection dynamics during the context of infection. P. aeruginosa is a gram-negative bacterium that commonly infects immunocompromised hosts. Recently, observations revealed that P. aeruginosa traffics to the gallbladder where it rapidly replicates. Current projects seek to uncover novel genetic elements required for replication in the gallbladder and understand the role that this organ plays in disease outcome and bacterial transmission.
- We investigate the development of multidrug resistance phenotypes in clinically relevant pathogens including Pseudomonas aeruginosa and Enterococcus faecium. We are currently exploring novel pathways involved in colistin resistance in P. aeruginosa and characterizing novel mutations in beta-lactamases that lead to antimicrobial resistance. Finally, we’ve begun a collaboration with the NMH clinical microbiology laboratory to apply whole-genome sequencing and molecular epidemiology to track outbreaks of vancomycin-resistant Enterococcus in the hospital.
Overall, our studies utilize a broad range of techniques including animal modeling, molecular and biochemical techniques, bacterial whole genome sequencing and antimicrobial resistance testing to explore bacterial pathogenesis and antimicrobial resistance. We hope that these basic insights will lead to improved diagnostics and therapeutics for bacterial diseases.
For more information, see Dr. Bachta's faculty profile.
Publications
View Dr. Bachta's complete list of publications in PubMed.
Contact Us
Kelly Bachta, MD, PhD at 312-503-3354

Vadim Backman
Professor, Medicine (Hematology and Oncology), Biochemistry and Molecular Genetics
Professor, McCormick School of Engineering
Elucidation of the function of the human genome and its dysregulation in disease

Vadim Backman
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Genetics, Epigenetics and Genomics
Prof. Backman’s lab is pioneering an exciting new field called physical genomics which focuses on studying the three-dimensional structure and dynamics of the genome and developing tools for its physical manipulation, with a goal of enabling reversible control of gene transcription patterns and the reprogramming of cells as a radically new approach to medicine and bioengineering. Breakthrough research from Backman’s Center for Physical Genomics and Engineering has recently demonstrated that the human genome physically functions as a self-assembling, reinforcement learning computational device - one that can read, write and store new cellular transcriptional memories. Through a convergence of novel super-resolution microscopy, computational modeling, genome mapping, AI and wet-lab bioengineering techniques, Backman’s group is developing the ability to rationally design genome folding states to control cellular behavior and function with implications for diseases such as cancer and Alzheimer’s, tissue regeneration, resiliency, human longevity and healthspan, and more.
For more information, see Dr. Backman's faculty profile or the Backman lab website.
Publications
View Dr. Backman's complete list of publications in PubMed.
Contact Us

Irina V Balyasnikova
Professor, Neurological Surgery
Advancing and developing translational targeted therapies for the treatment of glioblastoma and other brain malignancies

Irina V Balyasnikova
Research Area(s): Cancer Biology; Immunology; Neurobiology
The Balyasnikova Lab primarily focuses on developing novel targeted and immunotherapeutic approaches for treating malignant brain tumors such as GBM and metastases. In particular, Dr. Balyasnikova is interested in developing and utilizing antibodies and genetically engineered variants against tumor-specific antigens to enhance and deliver therapeutics specifically to the brain and other solid tumors. She also actively employs imaging technologies to understand the therapeutic response of malignant brain tumors to immunotherapy. Finally, as we look towards the future of brain cancer treatment, we are committed to the training of aspiring research professionals. For more information, please visit .
For more information, see Dr. Balyasnikova's faculty profile or the Balyasnikova Lab webpage
Publications
View Dr. Balyasnikova's complete list of publications in PubMed.
Contact Us

Grant D Barish
Associate Professor, Medicine (Endocrinology, Metabolism and Molecular Medicine)
Transcriptional regulators of inflammation and metabolism

Grant D Barish
Research Area(s): Cardiovascular and Renal Biology; Genetics, Epigenetics and Genomics; Immunology; Metabolism and Endocrinology; Reproductive Biology
Our laboratory has a longstanding interest in transcription factors and their roles in human disease. Over the past several years, we have identified novel functions for the germinal center transcriptional repressor BCL6 in regulating body fat distribution, insulin sensitivity, skeletal muscle mass, and metabolic dysfunction-associated steatotic liver disease. These studies continue to guide our efforts to define transcriptional pathways that may be therapeutically targeted in obesity-related metabolic disorders.
In parallel, we study CAMTA1-related disorder, a congenital cerebellar ataxia caused by monoallelic CAMTA1 mutations. Our work aims to define the role of CAMTA1 in neuronal gene regulation, determine how disease-associated mutations alter CAMTA1 function, and identify potential therapeutic strategies for affected individuals.
For more information, please see Dr. Barish's faculty profile.
Publications
See Dr. Barish's publications in PubMed.
Contact

Joseph T Bass
Professor, Medicine (Endocrinology, Metabolism and Molecular Medicine)
Professor, Weinberg College of Arts and Sciences
Circadian and metabolic gene networks in the development of diabetes and obesity

Joseph T Bass
Research Area(s): Cell Biology; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology
Circadian and metabolic gene networks in the development of diabetes and obesity
An epidemic of obesity and diabetes has continued to sweep through the industrialized world, already posing a risk to over one-third of the US population who are overweight or obese. Although both physical inactivity and overnutrition are tied to “diabesity,” recent evidence indicates that disruption of internal circadian clocks and sleep also play a role. The primary research focus in our laboratory is to apply genetic and biochemical approaches to understand the basic mechanisms through which the circadian clock regulates organismal metabolism. We anticipate that a better understanding of clock processes will lead to innovative therapeutics for a spectrum of diseases including diabetes, obesity, autoimmunity and cancer.
Studies of Clock Function in Beta Cell Failure and Metabolic Disease
Glucose homeostasis is a dynamic process subject to rhythmic variation throughout the day and night. Impaired glucose regulation leads to metabolic syndrome and diabetes mellitus, disorders that are also associated with sleep-wake disruption, although the molecular underpinnings of circadian glucose regulation have been unknown. Work from our laboratory first demonstrated an essential role of the intrinsic pancreatic clock in insulin secretion and diabetes mellitus and present efforts focus on dissecting the genomic and cell biologic link between clock function and beta cell failure (Nature, 2010, 2013).
Studies of Clock Regulation of Metabolic Epigenetics
In 2009 we first reported discovery that the circadian system plays a central role in metabolism through regulation of NAD+ biosynthesis (Science, 2009). NAD+ is a precursor of NADP+ and is required for macromolecule biosynthesis, in addition to functioning as an oxidoreductase carrier. NAD+ is also a required cofactor for the class III histone deacetylases (silencer of information regulators, SIRTs), nutrient-responsive epigenetic regulators Biochemical analyses show that SIRT1 deacetylates substrate proteins generating O-acetyl-ADP-ribose and nicotinamide, which is then regenerated to NAD+ by the enzyme nicotinamide phosphoribosyl transferase (NAMPT). We originally showed that CLOCK/BMAL1 directly control the transcription of Nampt and in turn control the activity of SIRT1—identifying a feedback loop composed of CLOCK/BMAL1-NAMPT/SIRT1. More recently, we have identified a role for the clock-NAD+ pathway in mitochondrial respiration (Science, 2013), and our present efforts include the analysis of clock-NAD+ regulation of cellular redox and epigenetic regulation, with the ultimate aim of applying such knowledge to studies of cell growth and stress response.
For more information, please see Dr. Bass' faculty profile or lab website.
Publications
See Dr. Bass' publications in PubMed.
Contact Info

Daniel Batlle
Professor, Medicine (Nephrology and Hypertension)
Renin angiotensin system in human diabetic kidney disease and rodent models of diabetic kidney disease and hypertension

Daniel Batlle
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Pharmacology
Dr. Batlle’s lab currently focuses on the renin angiotensin system as it relates to the understanding of this system in rodent kidney physiology. Of particular focus are the pathways and mechanisms that determine the enzymatic cleavage and degradation of Angiotensin II and other peptides within the system by ACE2-dependent and independent pathways. The lab uses a holistic approach involving ex vivo, in vitro and in vivo studies using various rodent models of diabetic and hypertensive kidney disease.
The lab is also involved in the search for biomarkers of kidney disease progression as part of the NIDDK Consortium on CKD. Other areas of research interest include nocturnal hypertension and the physiology and pathophysiology of electrolyte disorders such as distal renal tubular acidosis.
For more information, please see Dr. Batlle's faculty profile.
Publications
See Dr. Batlle's publications in PubMed.
Contact

Issam Ben-Sahra
Associate Professor, Biochemistry and Molecular Genetics
Decoding connections between signaling and metabolic networks

Issam Ben-Sahra
Research Area(s): Cell Biology; Metabolism and Endocrinology
The Ben-Sahra lab seeks to identify novel connections between oncogenic and physiological signals and cellular metabolism. My previous studies revealed new connections between mTORC1 (mechanistic Target of Rapamycin Complex I) signaling and de novo nucleotide synthesis pathways.
Using isotopic tracing experiments and genetic approaches, my lab investigates whether the additional signaling pathways such as PI3K/Akt, RAF/Erk, Hippo/Yap or AMPK could regulate metabolic pathways that supply small metabolites to sustain nucleotide synthesis independently of mTORC1 signaling. Furthermore, we are also interested in understanding how cells can sense changes in nucleotide levels. In addition to nucleotide metabolism, we also study connections between signaling pathway and global cancer cell metabolism. I predict that there could be points of regulations which could give selective advantages to cancer cells to grow and proliferate. The initial discovery that cancer cells exhibit atypical metabolic characteristics can be traced to the pioneering work of Otto Warburg, over the first half of the twentieth century.
Deciphering the interplay between oncogenic processes and metabolic pathways that contribute to metabolic reprogramming in a given setting may serve as a critical factor in determining therapeutic targets that yield greatest drug efficacy with marginal harmful effect on normal cells. Our research will enable further progress in the exploitation of unusual metabolic features in cancer as a means of therapeutic intervention.
For lab information and more, see Dr. Ben-Sahra's faculty profile and lab website.
Publications
See Dr. Ben-Sahra's publications on PubMed.
Contact
Contact Dr. Ben-Sahra.

Kynon J Benjamin
Assistant Professor, Psychiatry and Behavioral Sciences
Advanced computational genomics for the study of neurological disorders using disease-relevant models

Kynon J Benjamin
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Neurobiology
The HEART-GeN (Health Equity for Advancing Research and Technology using Genomic Neuroscience) research group is dedicated to improving neurotherapeutics for underrepresented communities by investigating the influence of genetic ancestry on molecular signatures in the brain. Our primary goals include uncovering how genetic ancestry impacts complex traits and enhancing our understanding of brain disorders across diverse genetic backgrounds. We employ an integrative approach that combines advanced computational tools (i.e., statistical genetics, machine learning, and deep learning) and disease-relevant models (i.e., postmortem brain tissues, brain organoids, and iPSC-derived glial cells) to provide comprehensive insights into brain function and disease mechanisms. Our research aims to elucidate the intricate interplay between genetic factors, epigenetic modifications, and environmental influences, ultimately striving to develop more effective and personalized neurotherapeutics. By addressing the critical gap in neuroscience research where certain populations have been historically underrepresented, HEART-GeN seeks to contribute to more inclusive and effective treatments for neurological disorders that account for the genetic diversity of human populations.
For lab information and more, see Dr. Benjamin's faculty profile.
Publications
See Dr. Benjamin's publications on PubMed.
Contact
Contact Dr. Benjamin.

M Cecilia Berin
Professor, Medicine (Allergy and Immunology)
Immune mechanisms of food allergy and tolerance

M Cecilia Berin
Research Area(s): Immunology
The Berin Lab is a translational immunology and allergy lab focused on the mechanisms of allergy and tolerance to foods. We study the contribution of allergen-specific T cells to the development, maintenance, and resolution of food allergy. This includes different manifestations of food allergy, including IgE-mediated and non-IgE-mediated food allergies. We also use samples from clinical trials to understand the immune basis of treatment response, our goal being the development of personalized medicine approaches as well as novel treatments for food allergy. We work with birth and infant cohorts to identify predictors of allergy, and study mechanisms of eczema and food allergy development with an emphasis on innate immune training.
The lab is heavily dependent on single cell approaches, primarily via flow cytometry but also transcriptional profiling, and primarily work with human specimens.
For more information, please see Dr. Berin's faculty profile.
Publications
See Dr. Berin's publications in PubMed.
Contact

Lisa R Beutler
Assistant Professor, Medicine (Endocrinology, Metabolism and Molecular Medicine)
Mechanisms of gut-brain communication underlying energy homeostasis

Lisa R Beutler
Research Area(s): Metabolism and Endocrinology; Neurobiology
Our goal is to understand how the gut and the brain communicate with each other to maintain body weight, and how this goes awry in diseases such as obesity. To accomplish this, we use a variety of techniques including optogenetics and calcium imaging in genetically modified mice. Projects in the lab focus both on how information about nutrients in the gastrointestinal tract are transmitted to the brain and how the brain regulates gastrointestinal tract function to optimize digestion and metabolism.
For more information, please see Dr. Beutler's faculty profile.
Publications
See Dr. Beutler's publications in PubMed.
Contact

Katherine Borden
Professor, Pharmacology
Disrupted RNA metabolism in cancer

Katherine Borden
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics; Pharmacology
Our lab’s work focuses on understanding the mechanisms and impact of dysregulated RNA metabolism in cancer using Acute Myeloid Leukemia as an example. RNA metabolism can be hijacked by cancer cells to re-write and/or amplify the message sent from the DNA. This in turn can underpin the oncogenic phenotype. We unearthed novel modes by which cancer cells hijack genetic information, in the form of messenger RNA, and demonstrated new mechanisms by which RNA maturation and RNA export influence cell physiology and support cancer growth. To do this, we use the eukaryotic translation initiation factor eIF4E as a model systems and demonstrated that it could act in splicing, RNA export as well as in translation of specific RNAs which in turn drives oncogenic pathways supporting cell motility, survival and other oncogenic properties. These findings drove first-in-class clinical trials demonstrating that targeting eIF4E and relevant RNA processing events represents a highly promising avenue for the development of novel therapeutics. These studies also revealed novel modes of drug resistance in patients that are targetable.
Note to students:
Our lab’s work focuses on understanding the mechanisms and impact of dysregulated RNA metabolism in Acute Myeloid Leukemia. RNA metabolism can be hijacked by cancer cells to re-write and/or amplify the message sent from the DNA. This in turn can underpin the oncogenic phenotype. Using the eukaryotic translation initiation factor eIF4E as a model, we unearthed novel modes by which cancer cells hijack genetic information, in the form of messenger RNA, and demonstrated new mechanisms by which RNA maturation and RNA export influence cell physiology and support cancer growth with a focus on AML cell motility and its interaction with bone marrow stroma. We conduct biochemical studies into the factors that imbue RNAs with eIF4E sensitivity at the levels of splicing, RNA export and translation. Another project involves defining the role of eIF4E phosphorylation in its nuclear functions in splicing and stimulation of capping and to define its importance to cell motility. We found that eIF4E drives programs to alter extracellular vesicle cargoes and seek to better define these activities and their impact on eIF4E-dependent motility. Many other related projects are also current interests in the lab.
Publications
View Dr. Borden's publications on PubMed.
For more information, please see Dr. Borden's faculty profile
Contact Us

Daniel J Brat
Professor, Pathology (Experimental Pathology), Pathology (Neuropathology)
Mechanisms underlying glioblastoma progression and regulators of asymmetric cellular division in glioblastoma stem cells

Daniel J Brat
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Neurobiology
Mechanisms Underlying Glioblastoma Progression
We investigate mechanisms of progression to glioblastoma (GBM), the highest grade astrocytoma, including genetics, hypoxia, and angiogenesis. Progression is characterized by tumor necrosis, severe hypoxia and microvascular hyperplasia, a type of angiogenesis. We propose that vaso-occlusion and intravascular thrombosis within a high grade glioma results in hypoxia, necrosis and hypoxia-induced microvascular hyperplasia in the tumor periphery, leading to neoplastic expansion outward. Since the pro-thrombotic protein tissue factor is upregulated in gliomas, we investigate mechanisms of increased expression and pro-coagulant effects.
In Silico Brain Tumor Research
We initiated an In Silico Center for Brain Tumor Research to investigate the molecular correlates of pathologic, radiologic and clinical features of gliomas using pre-existing databases, including as TCGA and Rembrandt. Using datasets and image analysis algorithms, we study whether elements of the tumor micro-environment, such as tumor necrosis, angiogenesis, inflammatory infiltrates and thrombosis, may correlate with gene expression subtypes in TCGA gliomas. We also have demonstrated the clinical relevance of TCGA subclasses within the lower grade gliomas using the Rembrandt dataset.
Regulators of Asymmetric Cellular Division in Glioblastoma Stem Cells
We study mechanisms that confer specialized biologic properties to glioma stem cells (GSC) in GBM. The Drosophila brain tumor (brat) gene normally regulates asymmetric cellular division and neural progenitor differentiation in the CNS of flies and, when mutated, leads to a massive brain containing only neuroblastic cells with tumor-like properties. We study the human homolog of Drosophila brat, Trim3, for its role in regulating asymmetric cell division and stem-like properties in GSCs. Trim3 may elicit its effects is through repression of c-Myc.
For more information, visit the faculty profile of Daniel Brat, MD, PhD or the Brat Lab website.
Publications
See Dr. Brat's publications in PubMed.
Contact

Melissa A Brown
Professor Emeritus, Microbiology-Immunology
Mechanisms underlying sex-related differences in autoimmune disease; meningeal inflammation and how it impacts CNS degenerative disease

Melissa A Brown
Research Area(s): Immunology; Neurobiology
Mast cells are found in most tissues including the gastrointestinal tract, respiratory tract, pancreas, synovium, brain, spinal cord and the secondary lymphoid organs. Best studied in the context of allergic disease, the widespread location of mast cells, the plethora of inflammatory mediators they produce and their ability to directly interact with T and B cells made them good candidates for exacerbating the inflammation associated with autoimmune diseases such as diabetes, arthritis and multiple sclerosis (MS).
We have utilized KitW/Wv mice, which are mast cell deficient, to study the contribution of these cells in a rodent model of MS, Experimental autoimmune/allergic encephalomyelitis (EAE). EAE is characterized by the T cell mediated orchestration that damages myelin and myelin producing cells in the CNS leading to severe neurological deficits due to loss of normal nerve conduction. We have shown that mast cells in the meninges are activated early in this disease and promote the opening of the blood brain barrier (BBB), vasculature that is relatively impermeable and normally sequesters the CNS from the entry of inflammatory cells. Our current studies focus on understanding how these mast cells influence these events. In the process, we have established a new paradigm for the mast cell mediated inflammation of the meninges in immunity and believe this information will likely impact the understanding of other CNS diseases.
A second line of research investigates the development of mast cells from early myeloid precursors. Mast cells share a common precursor with a related cell type, basophils. While mast cells are resident in tissues and their numbers remain relatively stable, basophils are induced in high numbers in the blood only in certain infection settings. We have demonstrated the Ikaros, a transcription factor, is essential for proper mast cell development. In Ikaros deficient mice, mast cell development is aberrant and basophils predominate in the absence of inflammatory signals. We are studying the events that underlie mast cell basophil-lineage choice in development by examining the molecular targets of Ikaros and its mode of action under basal and infection conditions.
For lab information and more, see Dr. Brown’s faculty profile and lab website.
Publications
See Dr. Brown's publications on PubMed.
Contact
Contact Dr. Brown at 312-503-0108 or the lab at 312-503-1013.

GR Scott Budinger
Professor, Medicine (Pulmonary and Critical Care), Cell and Developmental Biology
Mechanisms of aging and proteostasis stress

GR Scott Budinger
Research Area(s): Biochemistry and Structural Biology; Lung Biology; Microbiology
The Budinger lab studies the mechanisms underlying the loss of organismal resilience during aging, focusing on the hypothesis that some of these changes are induced by chronic stress to the proteostasis network. We are particularly interested in how proteostasis stress during aging induces dysfunction in tissue resident macrophages in the lung, brain and skeletal muscle that are important for organ repair. We use pneumonia as a model of systemic organismal stress in animals that mimics many of the features we see in patients hospitalized for pneumonia in our hospital.
For lab information and more, see Dr. Budinger’s faculty profile.
Publications
See Dr. Budinger's publications on PubMed.
Contact
Contact Dr. Budinger or the administrative office at 312-908-7737.
Irina V Budunova
Professor, Dermatology, Urology
Role of the glucocorticoid receptor in carcinogenesis and stem cell maintenance, development GR-targeted therapies in skin
Irina V Budunova
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology; Immunology; Pharmacology
The current projects in Dr. Budunova’s lab are centered on the role of the glucocorticoid receptor (GR) as a tumor suppressor gene in skin. We showed that skin-specific GR transgenic animals are resistant to skin carcinogenesis and GR KO animals are more sensitive to skin tumor development. We are also interested in the role of GR in the maintenance of skin stem cells (SC). We found that GR/glucocorticoids inhibit the expression of numerous SC markers in skin including CD34- a marker of hair follicular epithelial SC and reduce the proliferative potential of skin SCs.
The glucocorticoids remain among the most effective and frequently used anti-inflammatory drugs in dermatology. Unfortunately, patients chronically treated with topical glucocorticoids, develop side effects including cutaneous atrophy. GR controls gene expression via (i) transactivation that requires GR dimerization and binding as homo-dimer to gene promoters and (ii) transrepression that is chiefly mediated via negative interaction between GR and other transcription factors including pro-inflammatory factor NF-kB. In general, GR transrepression is the leading mechanism of glucocorticoid anti-inflammatory effects, while many adverse effects of glucocorticoids are driven by GR transactivation.
Our laboratory has been involved in delineation of mechanisms underlying side effects of glucocorticoids in skin. Using GRdim knockin mice characterized by impaired GR dimerization and activation, we found that GR transactivation plays an important role in skin atrophy. These data suggested that non-steroidal selective GR activators (SEGRA) that do not support GR dimerization, could preserve therapeutic potential of classical glucocorticoids but have reduced adverse effects in skin. We are testing effects of the novel SEGRA called Compound A– a synthetic analog of natural aziridine precursor from African bush Salsola Botch in skin. We have also established anti-cancer GR-dependent activity of Compound A in epithelial and lymphoma cells.
Using knockout mice for the major GR target genes including Fkbp5 (GR chaperone) and DDIT4/REDD1 (one of the major negative regulators of mTORC), we discovered that blockage of Fkbp5 and REDD1 significantly changes GR function and greatly protects skin against glucocorticoid-induced atrophy. This suggests a novel GR-targeted anti-inflammatory therapy where glucocorticoids are combined with inhibitors of GR target genes.
For more information, please see Dr. Budunova’s faculty profile.
Publications
See Dr. Budunova's publications in PubMed.
Contact Budunova Lab
Contact the Budunova Lab at 312-503-4669 or visit in the Montgomery Ward Building, 303 E. Chicago Avenue, Ward 9-015, Chicago, IL 60611

Serdar E Bulun
Professor, Obstetrics and Gynecology (Reproductive Science in Medicine), Obstetrics and Gynecology (Reproductive Endo and Infertility)
Estrogen metabolism in breast cancer, endometriosis and uterine fibroids

Serdar E Bulun
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics; Reproductive Biology
The laboratory research of Serdar E. Bulun, MD, focuses on studying estrogen biosynthesis and metabolism, in particular aromatase expression, in hormone-dependent human diseases such as breast cancer, endometriosis and uterine fibroids.
A team of investigators works on understanding the epithelial-stromal interactions and aromatase overexpression in breast cancer tissue. Because aromatase inhibitors treat breast tumors primarily via suppressing intratumoral estrogen biosynthesis, these efforts are important for discovering new targets of treatment.
Another team studies endometriosis. Basic data from this laboratory led to the introduction of aromatase inhibitors into endometriosis treatment. Human tissues and a primate model are used to elucidate cellular and molecular mechanisms responsible for the development of endometriosis.
Regulation of aromatase expression is also studied in uterine fibroids, benign tumors that are dependent on estrogen for growth, by a third team.
A fourth team is investigating the link between progesterone action and estrogen inactivation in normal endometrium and endometriosis.
Lastly, a fifth team has identified novel mutations that cause familial excessive estrogen formation syndrome. This syndrome is characterized by short stature, gynecomastia and hypogonadism in males and early breast development and irregular menses in females. In this syndrome, heterozygous inversions in chromosome 15q21, which cause the coding region of the aromatase gene to lie adjacent to constitutively active cryptic promoters that normally transcribe other genes, result in estrogen excess owing to the overexpression of aromatase in many tissues.
For more information, please see Dr. Bulun's faculty profile.
Publications
See Dr. Bulun's publications in PubMed.
Contact

Paul W Burridge
Associate Professor, Pharmacology
Application of human induced pluripotent stem cells to study the pharmacogenomics of chemotherapy off-target toxicity and efficacy

Paul W Burridge
Research Area(s): Cancer Biology; Cardiovascular and Renal Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Pharmacology

Research Description
The Burridge Lab studies the role of the genome in influencing drug responses, known as pharmacogenomics or personalized medicine. Our major model is human induced pluripotent stem cells (hiPSC), generated from patient's blood or skin. We use a combination of next generation sequencing, automation and robotics, high-throughput drug screening, high-content imaging, tissue engineering, electrophysiological and physiological testing to better understand the mechanisms of drug response and action.
Our major effort has been related to patient-specific responses to chemotherapy agents. We ask the question: what is the genetic reason why some patients have a minimal side effects to their cancer treatment, whilst others have encounter highly detrimental side-effects? These side-effects can include cardiomyopathy (heart failure or arrhythmias), peripheral neuropathy or hepatotoxicity (liver failure). It is our aim to add to risk-based screening by functionally validating genetic changes that predispose a patient to a specific drug response.
Recent Findings
- Human induced pluripotent stem cells predict breast cancer patients’ predilection to doxorubicin-induced cardiotoxicity
- Chemically defined generation of human cardiomyocytes
Current Projects
- Modeling the role of the genome in doxorubicin-induced cardiotoxicity using hiPSC
- Investigating the pharmacogenomics of tyrosine kinase inhibitor cardiotoxicity
- hiPSC reprogramming, culture and differentiation techniques
- High-throughput and high-content methodologies in hiPSC-based screening
For lab information and more, see Dr. Burridge’s faculty profile and lab website.
Publications
See Dr. Burridge's publications on PubMed.
Contact
Contact Dr. Burridge at 312-503-4895.

Qi Cao
Professor, Urology
Epigenetic regulation in prostate cancer progression

Qi Cao
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics
Research Description
For two decades, my work has sat at the intersection of epigenetics and cancer biology, resulting in over 70 peer-reviewed publications. Our research primarily explores how histone and RNA modifications, alongside non-coding RNAs, drive the progression of breast and prostate cancers. Key milestones of our discoveries include identifying EZH2 as a critical biomarker for aggressive malignancy and discovering the genomic loss of miR-101. Notably, my laboratory was the first to describe the non-canonical roles of BMI1 and EZH2—demonstrating their PRC-independent influence on AR protein stability and IRES-driven translation. By establishing these proteins as essential therapeutic targets, my research has provided the mechanistic foundation for the development of EZH2 and BMI1 inhibitors currently being pursued by several pharmaceutical companies.
For more information, see Dr. Cao's faculty profile.
Publications
See Dr. Cao's publications on PubMed.
Contact
Contact Dr. Cao.

Gabriela Caraveo Piso
Assistant Professor, Neurology (Movement Disorders), Pharmacology
Role of Ca2+ signaling in synucleinopathies using diverse model systems from yeast to human neurons

Gabriela Caraveo Piso
Research Area(s): Cell Biology; Neurobiology
We are focused on a group of neurodegenerative diseases collectively known as synucleinopathies, characterized by the aggregation of α-synuclein (α-syn). These include Parkinson's Disease, Dementia with Lewy bodies and Multiple Systems Atrophy among others. We use diverse systems that span from yeast to mammalian models to study these diseases. In particular, we are interested in the role Ca2+ signaling plays in the toxicity caused by α-syn and to delineate basic mechanisms of Ca2+ signaling relevant to neuronal physiology.
For more information, visit Dr. Caraveo Piso's faculty profile or the Caraveo Piso Lab website.
Publications
Please see Dr. Caraveo Piso's publications on PubMed.
Contact Information
Assistant Professor in Neurology
Ward 10-150
312-503-4492

Gemma L Carvill
Associate Professor, Neurology (Epilepsy/Clinical Neurophysiology), Pharmacology, Pediatrics
Genetic causes and pathogenic mechanisms that underlie epilepsy.

Gemma L Carvill
Research Area(s): Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology
The primary goal of our research is to use gene discovery and molecular biology approaches to identify new treatments for epilepsy. We aim to 1) identify the genetic causes of epilepsy, 2) use stem cell models to understand how genetic mutations can cause epilepsy, 3) develop and test new therapeutics for this condition. Our work is based on the promise of precision medicine where knowledge of an individual’s genetic makeup shapes a personalized approach to care and management of epilepsy.
Current Projects
- Next generation sequencing in patients with epilepsy
- Alternative exon usage during neuronal development
- Identify the regulatory elements that control expression of known epilepsy genes
- Stem cell genetic models for studying the epigenetic basis of epilepsy
For more information, see Dr. Carvill's faculty profile or the Carvill Lab Website.
Publications
Please see Dr. Carvill's publications on PubMed.
Contact Information

Debabrata Chakravarti
Professor, Obstetrics and Gynecology (Reproductive Science in Medicine), Pharmacology
Epigenome and 3D chromatin organization dysregulations define human cancers and reproductive diseases

Debabrata Chakravarti
Research Area(s): Cancer Biology; Cell Biology; Genetics, Epigenetics and Genomics; Pharmacology; Reproductive Biology
Dr. Chakravarti’s research is focused on understanding epigenetic and transcriptional regulation of human tumorigenesis. One of his research projects is focused on understanding the mechanisms that drive the development of uterine fibroids and endometriosis that affect an alarmingly high number of all women. In another project, Dr. Chakravarti’s research team investigates molecular underpinning of contribution of transcription factors, cofactors and epigenomic and 3D genome reorganization regulation of prostate Cancer that affects a large number of men worldwide. In a third project the laboratory determines the role of protein cofactors in regulation of cell cycle genes. Thus, our work interfaces both fundamental and translational research on diseases that affect humankind. It is our hope that when combined with results from others, our research will contribute to the development of future therapeutics. Dr. Chakravarti gratefully acknowledges continuous funding support from the NIH and key roles of his lab members and collaborators in the overall success of the Chakravarti Laboratory.
Dr. Chakravarti also enjoys teaching. He has continuously taught both medical and graduate students. He serves on numerous Ph.D thesis committees. He has trained a large number of graduate students and postdoctoral fellows some of whom are now independent investigators at this and other institutions.
For more information, please see, visit the Dr. Chakravarti's faculty profile.
Publications
See Dr. Chakravarti's publications in PubMed.
Associate Editor: Endocrinology 2017-present; Editorial Board: Molecular Endocrinology 2011- present, Mol. Cell. Biol. 2014-present
The Editor of a Book volume on “Regulatory Mechanisms in Transcriptional Signaling” in Progress in Molecular Biology and Translational Science (Vol 87), published in Aug 2009, Academic Press, Chakravarti, D. Editor
Contact Us
312-503-1641

Navdeep S Chandel
Professor, Medicine (Pulmonary and Critical Care), Biochemistry and Molecular Genetics
Mitochondria as a signaling organelle; reactive oxygen species as the primary signal for metabolic adaptation, differentiation and proliferation

Navdeep S Chandel
Research Area(s): Cancer Biology; Cell Biology; Immunology; Metabolism and Endocrinology; Microbiology; Neurobiology
Research Goals
For decades, mitochondria have been primarily viewed as biosynthetic and bioenergetic organelles generating metabolites for the production of macromolecules and ATP, respectively. Dr. Chandel’s work has elucidated that mitochondria have a third distinct role whereby they participate in cellular signaling processes through the release of reactive oxygen species (ROS) and metabolites independent of ATP and macromolecule production. Our work has implicated the necessity of mitochondrial ROS for multiple biological processes including hypoxic activation of HIFs, cellular differentiation, and adaptive immunity. Previously, the dogma in the field had been that mitochondrial ROS are only produced in pathological settings to cause both lipid, protein and DNA damage. However, our work demonstrates that mitochondrial ROS are utilized as messengers to maintain normal biological and physiological functions. Our studies suggest that the current widespread use of antioxidants is likely to be detrimental rather than beneficial for alleviating a myriad of diseases as this could interfere with normal physiological processes. Recently, our work has shown that mitochondria release the metabolite L-2HG, which increases histone and DNA methylation to control hematopoietic stem cell (HSC) differentiation and regulatory T cell (Treg) function, respectively. In summary, our lab has been instrumental in changing our view of mitochondria from the “powerhouses" of cell to “signaling organelles”. He has trained past/present 27 graduate students.
Current Projects
Cancer Metabolism:
In the field of cancer, our work was the first to genetically demonstrate the necessity of mitochondrial respiratory chain is necessary for tumor growth and angiogenesis in vivo. Our recent work points to key role of the respiratory chain in supporting biomass of cancer cells needed for in vivo tumor growth. These findings were seminal for the cancer metabolism field since the 1920s the prevailing idea was that only increased aerobic glycolysis (i.e. the Warburg Effect) was the dominant metabolic reprogramming event in cancer cells and endothelial cells. Another key finding in the lab was genetically elucidating that the anti-diabetic drug metformin prevents tumorigenesis by inhibiting mitochondrial complex I within cancer cells. Finally, our work provided a conceptual model whereby mitochondria generated ROS can trigger local signaling events in the cells to promote growth but can be elevated to levels that are detrimental. Thus, tumor cells increase their antioxidant capacity to prevent ROS induced cell death. Our current projects involve the use of CRISPR based genetic screens to uncover metabolic vulnerabilities in the presence of standard care of therapy.
Viral Pneumonia:
Viral pneumonia is currently among the most common causes of death in the world. Viral pneumonia impairs tissue repair leading to both in-hospital death and prolonged hospital-acquired disability. Two leading causes of viral pneumonia are influenza A virus and SARS-CoV-2 virus. We use mouse models to test whether metabolism can modulate inflammation, host defense and repair to the lungs upon exposure to influenza A virus.
Aging:
Metformin is a biguanide drug that boosts healthspan in multiple model organisms and humans. Metformin in humans is used as the first-line therapy for type II diabetes. In addition, metformin has anti-tumor and anti-inflammatory effects which likely contribute to its geroprotective activity. Currently, the TAME trial will test whether metformin can delay age-related chronic diseases in humans. However, metformin’s the direct mechanism of action is poorly understood. The mechanism of metformin action must be considered at two levels: (1) the molecular target of metformin and (2) the therapeutic pathways triggered by metformin acting on its molecular target in different cell types. A leading hypothesis is that metformin inhibits mitochondrial complex I (NADH dehydrogenase) to exert anti-aging effects. Currently, we are using genetic models to determine the essentiality of mitochondrial complex I inhibition in gut epithelium, hepatocytes, and immune cells for metformin to exert its anti-diabetic and anti-inflammatory effects.
Neurological diseases:
Neurological diseases including amyotrophic lateral sclerosis (ALS), Parkinson’s disease (PD), and Alzheimer disease (AD) have all been linked to metabolic dysregulation. We are using mouse models and IPSCs to test whether metabolic dysfunction is a causal agent rather than a hallmark of these diseases. This work is complemented by our work on primary mitochondrial disease mouse models that manifest neurological symptoms.
Publications
See Dr. Chandel's publications in PubMed.
For more information, please see Dr. Chandel's faculty profile or visit the Chandel Lab website.
Contact Us
Contact Dr. Chandel’s Lab at 312-503-1792

Shi-Yuan Cheng
Professor, Neurology (Neuro-oncology)
Cancer stem cell biology, cellular signaling and therapy responses in human brain tumors, particularly glioblastoma (GBM)

Shi-Yuan Cheng
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Neurobiology
Our lab broadly studies cancer stem cell biology, cellular signaling, RNA biology, and therapy responses in human brain tumors, in particular, glioblastoma (GBM). We have a range of different projects currently underway in glioma cell lines, gliomas stem-like cells (GSCs), patient-derived xenograft (PDX) GBM model, human iPSC-derived glioma organoid model, orthotopic glioma xenograft model in mice, and clinical glioma tumor specimens. Our current research focuses on novel mechanisms/cellular signaling of GSC biology, tumorigenesis, progression, and therapy responses of GSCs and GBMs.
Roles of RNA alternative splicing and RNA-binding proteins in glioma
RNA alternative splicing (AS), an evolutionarily conserved co-transcriptional process, is an important and influential determinant of transcriptome and proteome landscapes in normal and disease states such as cancer. AS is regulated by a group of RNA binding proteins (RBPs) that bind to the cis-acting elements in proximity to a splice site thus affecting spliceosome assembly. In cancers, altered expression of or mutations in RBPs result in dysregulated AS that impacts cancer biologic properties. We have established AS/RBP networks that are dysregulated in both adult and pediatric gliomas through bioinformatic analysis of both public and our own datasets of clinical glioma tumors. We are investigating the biological significance of AS/RBPs dysregulation in glioma progression and therapy response by using human iPSC-derived glioma organoid model and GSC brain xenograft models in animals. In addition, we are exploring novel therapeutic approaches of targeting glioma-associated AS/RBP networks to treat GBMs.
Roles of Non-coding RNAs in glioma
Non-coding RNAs (ncRNAs), including long non-coding RNAs (lncRNAs) and circular RNAs (circRNAs), act as transcription repressors or inducers of gene expression or functional modulators in all multicellular organisms. Dysregulated ncRNAs plays critical roles in cancer initiation, progression and responses to therapy. We study the mechanisms by which deregulated expression of lncRNAs or circRNAs influence GBM malignant phenotypes through interactions with signaling pathways. We study the molecular consequences and explore clinical applications of modulating ncRNAs and related oncogenic signaling pathways in GBM. We are establishing profiles of ncRNAs in clinical gliomas and patient-derived GSCs, and study mechanisms and biological influences of these ncRNAs in regulating GSC biology and GBM phenotypes.
Aberrant DNA and RNA structures in therapy-resistant GBM
Standard of care treatment for GBM includes the DNA damaging agent temozolomide (TMZ), which has a known mechanism of action to target and mutate guanine bases. With this knowledge in hand, we sought to determine the effects of guanine (G) mutations in DNA and RNA secondary structure. G’s are important for creating structures like g-quadruplexes in both DNA and RNA which can affect changes in translation or be used as docking sites for DNA repair and RNA binding proteins. Using whole genome sequencing data along with isogenic drug sensitive and resistant lines, we are investigating the role of G mutations in DNA and RNA secondary structure to determine potential therapeutic avenues with the help of a chemical biologist to create novel drugs to target these TMZ-induced aberrant pathways.
Targeting autophagy to treat glioma
Autophagy is an evolutionarily conserved process that removes unnecessary or dysfunctional components through a lysosome-dependent regulated mechanism, thus serving as a protective mechanism against stressors and diverse pathologies including cancer. We study mechanisms by which phosphorylation, acetylation and ubiquitination of autophagy-related proteins regulate GSC and GBM phenotypes and autophagic response, which, in turn contributes to tumor cell survival, growth and resistance to therapy. We investigate whether disruption of these post-translational processes in autophagy-related proteins inhibits autophagy and enhances the efficacy of combination therapies in GBMs. In collaboration with a medicinal chemist, we are characterizing a next generation of novel autophagy inhibitors that specifically target a key autophagy regulator that we recently reported.
Multi-omics and GBM non-responsiveness to immunotherapies
GBM is categorized as a “cold” tumor that does not respond to current immunotherapies using various immune-checkpoint blockers. Although extensive efforts have been made to sensitize GBM to immunotherapies, the mechanistic studies to determine alternative therapies from understanding the underlying signaling and clinical trial results are still disappointing. We are interested in utilizing the information of multi-omics of clinical gliomas, in particular, proteomics profiling in relation to genomic and epigenomic profiling, to identify potential protein targets that could be the major modulators through post-translational modifications in these “cold” GBM tumors. We will also consider the involvement of tumor microenvironment and immune cells in these conditions. These studies are a brand-new direction that are high-risk and high-reward to turn “cold” GBM tumors to immunotherapy responsive tumors.
For more information, please see Dr. Cheng's faculty profile and lab website.
Publications
View Dr. Cheng's complete list of publications in PubMed.
Contact Us
Shi-Yuan Cheng, PhD at 312-503-5314
Visit us on campus in the Lurie Building, Room 6-119, 303 E Superior Street, Chicago, Illinois 60611.

Rex L Chisholm
Professor, Cell and Developmental Biology, Surgery
Molecular motors and cell motility

Rex L Chisholm
Research Area(s): Cell Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics
Movement is a fundamental characteristic of life. Cell movement is critical for normal embryogenesis, tissue formation, wound healing and defense against infection. It is also an important factor in diseases such as cancer metastasis and birth defects. Movement of components within cells is necessary for mitosis, hormone secretion, phagocytosis and endocytosis. Molecular motors that move along microfilaments (myosin) and microtubules (dynein) power these movements. Our goal is to understand how these motors produce movement and are regulated. We wish to define their involvement in intracellular, cellular and tissue function and disease—with the long-term goal of developing therapies for the treatment of diseases caused by defects in these molecular motors.
Our work involves the manipulation of myosin and dynein function in the single celled eukaryote Dictyostelium, cultured mammalian cells and transgenic and knockout mice. Yeast two-hybrid screens to identify proteins that interact with or regulate myosin and dynein and characterization of gene expression are being used to define the pathways regulating myosin and dynein. To analyze the biological significance of myosin and dynein, we use confocal and digital microscopy of living cells, analysis of cell movement, vesicle transport and cell division. We employ biochemical techniques including heterologous expression, enzyme purification and characterization and analysis of how phosphorylation state affects physiological function. We are pursuing signal transduction studies to understand the physiologically important pathways that regulate cell motility and biophysical studies such as in vitro motility assays to understand how these molecular motors function at the molecular level.
For lab information and more, see Dr. Chisholm's faculty profile.
Publications
See Dr. Chisholm's publications on PubMed.
Contact
Contact Dr. Chisholm at 312-503-3209.

Insup Choi
Assistant Professor, Pathology
Roles of Immune Cell Senescence in Neurodegenerative Disease Progression

Insup Choi
Research Area(s): Cell Biology; Immunology; Neurobiology
Understanding the neuronal environment is essential for addressing neurodegenerative diseases. Microglia, as brain-resident macrophages, play a crucial role in this context. My research focuses on senescent microglia in Parkinson's and Alzheimer's diseases. We use genetic mouse models and cell culture systems to explore how disease conditions promote microglial senescence, the mechanisms linking these senescent cells to neurotoxicity, and the role of genetic risk factors. By employing reporter mice to detect senescent cells, we aim to uncover insights into microglial senescence and its implications for neurodegenerative diseases, potentially revealing new therapeutic targets.For further information, please also see Dr. Choi's faculty profile.
Publications
See Dr. Choi's publications on PubMed.
Contact
Contact Dr. Choi.

Nicholas Cianciotto
Professor, Microbiology-Immunology
Pathogenesis of Legionella pneumophila and Stenotrophomonas maltophilia

Nicholas Cianciotto
Research Area(s): Immunology; Microbiology
L. pneumophila, the agent of Legionnaires' disease, is a classic environmental, opportunistic pathogen. The aim of our research is to characterize the bacterial genes and gene products that promote the occurrence of Legionnaires' disease.
S. maltophilia is an environmental gram-negative bacterium that is being increasingly associated with an array of human infections, including most notably pneumonia. The emergence of S. maltophilia as a significant health concern is due in part to its marked antibiotic resistance. Our lab has developed murine models of lung infection and is currently identifying virulence factors produced by this important new pathogen.
We employ a multi-faceted approach toward understanding the pathogenesis and natural history of bacterial infectious disease, with the hope that basic insights will lead to new methods of disease prevention, diagnosis, or treatment.
For lab information and more, see Dr. Cianciotto’s faculty profile and lab website.
Publications
See Dr. Cianciotto's publications on PubMed.
Contact
Contact Dr. Cianciotto at 312-503-0385 or the lab at 312-503-1034.

Yingzi Cong
Professor, Medicine (Gastroenterology and Hepatology), Microbiology-Immunology, Pathology
Immune regulation of host-microbiota at the intestine surface and the pathogenesis of inflammatory bowel diseases

Yingzi Cong
Research Area(s): Immunology; Metabolism and Endocrinology; Microbiology
The Cong lab focuses on the pathogenesis of inflammatory bowel diseases (IBD), including Crohn’s disease and ulcerative colitis, and the immune regulation of host-microbiome interaction at the intestine surface, which reaches 300m2 in humans and is exposed daily to large amounts of innocuous antigens from the diet and the resident microbiota/pathogens using various cutting-edge approaches, including genetic, epigenetic, metabolic, molecular biology, and immunological approaches, combining with the use of germ-free animals to dissect the host-microbiome networks which maintain intestinal homeostasis and regulate IBD and autoimmunity. Using state-of-the-art technologies, including single-cell RNA-seq, spatial expression, ATAC-seq, gut bacterial 16s/short-gun seq, and various unique transgenic/knockout mouse models, we have worked on mucosal T cells, innate cells, B cells, and intestinal epithelial cells and identified several specific genes and gut bacteria/bacterial metabolites critical for immune regulation and the pathogenesis of IBD. Our lab heavily collaborates with the IBD team at Northwestern Medicine for translational studies.
The ongoing research projects include:
- Gut bacterial metabolite regulation of mucosal immune responses and IBD.
- STING regulation of mucosal immune responses and IBD.
- Gut tissue memory T cell regulation of IBD.
- microRNA 10a regulation of T cell responses and IBD
- Intestinal epithelial cell-T cell crosstalk in response to gut microbiota.
- Intestinal IgA regulation of mucosal immune responses and the pathogenesis of IBD.
For more information, see Dr. Cong's faculty profile.
Publications
See Dr. Cong's publications

Anis Contractor
Professor, Neuroscience, Psychiatry and Behavioral Sciences
Professor, Weinberg College of Arts and Sciences
Link between synaptic dysfunction and neuropsychiatric disorders and neurodevelopmental disorders

Anis Contractor
Research Area(s): Neurobiology; Pharmacology
Research in our laboratory is directed at understanding the mechanisms of synaptic transmission and plasticity and the role that glutamate receptors have in brain function and pathology. We use a multidisciplinary approach including in vitro electrophysiological recording, optogenetics, cellular imaging, mouse behavior and biochemical techniques. We ultimately seek to understand the link between synaptic dysfunction and neuropsychiatric disorders and neurodevelopmental disorders. Current projects are investigating altered synaptic signaling in mouse models of obsessive compulsive disorder, schizophrenia and fragile X syndrome.
For lab information and more, see Dr. Contractor's faculty profile.
Publications
See Dr. Contractor's publications on PubMed.
Contact
Contact Dr. Contractor 312-503-1843 or the lab at 312-503-0276.

Lee A Cooper
Professor, Pathology (Experimental Pathology)
Computational and integrative pathology

Lee A Cooper
Research Area(s): Computational Biology and Bioinformatics
Our research spans the spectrum from application of novel computational methods to improve diagnosis and classification of disease to fundamental research in AI/ML methodology and software tool development.
The pathology lab is a rich resource of data that is undergoing significant transformation with the introduction of new imaging techniques and advances in genomics. Our goal is to create and apply the tools needed to interpret the data from multifaceted profiling of tissues to improve diagnostics and to provide basic science investigators with tools to generate new insights. To accomplish these goals we focus on:
- Developing new methodology to analyze whole-slide imaging, genomic, and spatially resolved molecular data.
- Creating software and infrastructure to aid clinical translation of advanced computational methods.
- Developing rich datasets that combine imaging and clinical data at scale using materials from the pathology lab.
As computational researchers we are not bound to a single disease, but we have worked in areas including breast cancer, hematologic malignancies, peri-natal pathology, and transplant. Example projects include:
- Prognosis from deep characterization of the breast cancer tumor microenvironment.
- Improving detection and grading of prostate cancer.
- Survival analysis methodology from high-dimensional multi-omic data.
- Improving classification of hematologic malignancies.
For more information, see Dr. Cooper's faculty profile or lab website.
Publications
See Dr. Cooper's publications

Carla Marie Cuda
Assistant Professor, Medicine (Rheumatology)
Cellular and molecular mechanisms underlying neuropsychiatric manifestations of systemic lupus erythematosus

Carla Marie Cuda
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Immunology; Neurobiology
My research program centers on uncovering mechanisms underlying neuropsychiatric manifestations of systemic lupus erythematosus (SLE) via basic and translational methodology. SLE is a chronic and multi-systemic autoimmune disease plaguing 1.5 million Americans. Among the many organ systems affected, symptoms associated with the nervous system (NP-SLE) may appear in up to 90% of SLE patients depending on the attribution model and be among the earliest signs of SLE. NP-SLE can manifest in seizures, strokes, movement disorders, mood disorder, anxiety disorder, acute confusional state, psychosis, acute encephalopathy and/or chronic neurocognitive dysfunction. Despite the devastating impact of NP-SLE on health-related quality-of-life, our understanding of causal mechanisms is limited.
Microglia, the resident innate immune cell of the brain, have recently been implicated in NP-SLE. Studies show histological evidence of microglia activation in NP-SLE patients and mouse models of disease. Microglia display regional, functional, and disease-associated heterogeneity; in particular, a disease-associated microglia (DAM) subset was recently identified in Alzheimer’s disease and aging. DAM co-localize with amyloid-b plaques, are enriched for genes involved in lysosomal, lipid metabolism and phagocytic pathways and may play a protective role in the early stages of AD by reducing amyloid-b plaque burden. However, very few studies have examined microglial heterogeneity or contributions to NP-SLE. Our group is the first to show that expression of a shared NP-SLE transcriptional signature and DAM-associated genes correlates with the severity of hippocampal- and cerebellar-associated behavioral deficits in microglia isolated from two NP-SLE models prior to overt systemic disease. Further, our single-cell RNA-sequencing (scRNA-seq) data identify DAM NP-SLE-prone mice. However, DAM in NP-SLE are enriched for genes associated with antigen presentation but depleted for genes associated with phagocytosis, which is in contrast to DAM in AD that are critical for amyloid-b plaque-associated phagocytic functions. We also find that restricted expression of the DAM transcriptional program in NP-SLE DAM corresponds to improved behavioral outcomes in NP-SLE-prone mice following treatment with fingolimod, a sphingosine-1-phosphate receptor modulator that reduces microglia activation and improves blood brain barrier integrity. These discoveries mark the first to implicate DAM as a potentially pathogenic microglia subset in NP-SLE, which is in contrast to their proposed protective role in early AD development.
We also find that increasing numbers of macrophages in the brain correlates with worsening severity of behavioral deficits early in life and an expanded brain macrophage subset in end-stage disease via scRNA-seq analysis that is detectable at two months of age. Since a worse prognosis is suggested when NP-SLE accompanies lupus nephritis and infiltrating macrophages contribute to organ-specific SLE pathogenesis, we hypothesize that macrophage-specific mechanisms underlying systemic disease are conserved in kidney and brain and detectable in circulating monocytes prior to extravasation into the affected end-organ.
Further, we are translating our findings to human disease by evaluating cerebrospinal fluid (CSF)-resident microglia-like cells and macrophages as well as circulating monocyte subsets from NP-SLE patients via scRNA-seq to interrogate the penetrance of causative mechanisms. Thus, we are interrogating microglial and macrophage heterogeneity to dissect cell subset-specific contributions to disease with the ultimate goal of identifying critical populations and/or pathways that may serve to improve diagnostic and/or therapeutic strategies in NP-SLE.
Publications
View lab publications via PubMed.
For more information, visit the faculty profile page of Carla Cuda, PhD.
Contact Us
Email Dr. Cuda

Weiguo Cui
Professor, Pathology (Experimental Pathology)
T cell differentiation and function in the context of infection and cancer

Weiguo Cui
Research Area(s): Cancer Biology; Immunology; Microbiology
The Cui Lab focuses on investigating the differentiation, function, and epigenetic regulation of T cells in diverse inflammatory contexts, including acute and chronic viral infections, autoimmune disorders, and cancer. T cells play a crucial role in the adaptive immune system and are essential for defending against bacteria, viruses, and malignant cells. Leveraging state-of-the-art technologies, such as single-cell RNA sequencing, TCR sequencing, ATAC sequencing, multi-color flow cytometry, and transgenic mouse models, our goal is to gain a comprehensive understanding of how T cells respond to different antigenic and inflammatory stimuli. We aim to unravel the mechanisms underlying their ability to control viral infections and combat cancer.
A major focus of our research is to dissect the cellular and molecular mechanisms that govern T cell functional exhaustion. We investigate the factors contributing to T cell exhaustion and aim to identify key regulators involved in this process. By understanding the underlying mechanisms, we strive to develop innovative approaches to reverse T cell exhaustion and enhance their functionality. This line of investigation holds great potential for developing novel therapeutic strategies to effectively treat chronic viral infections and combat cancer.
For more information, visit the faculty profile page of Weiguo Cui.
Publications
View Dr. Cui's publications via PubMed.
Contact Us
Email Dr. Cui

Richard T D'Aquila
Professor, Medicine (Infectious Diseases)
Pathogenesis of human immunodeficiency virus (HIV)

Richard T D'Aquila
Research Area(s): Immunology; Microbiology; Pharmacology
The D’Aquila laboratory studies the pathogenesis of human immunodeficiency virus (HIV) persistence/latency and mechanisms of host “intrinsic” immune control of HIV replication. The long-term goal is to develop innovative strategies for a “functional cure” of HIV. We are developing novel interventions to induce sustained remissions after defined-duration antiretroviral therapy (ART). This includes bench-to-bedside research to characterize / leverage / increase APOBEC3 (A3) “intrinsic” immune defenses against HIV, diminish T cell activation-fueled HIV replication and innovate strategies to minimize latency/persistence of HIV.
In papers published in recent years, the D’Aquila laboratory proved that APOBEC3G and F are active in producer and target cells against wild-type (Vif-positive) HIV-1 at higher physiological levels of expression. Evidence was published that higher levels of A3G protein and activity contribute to the multiple mechanisms of spontaneous control of HIV-1 without ART seen in rare “elite controllers”. The laboratory has also identified that limiting c-Myc-dependent effects of the host T cell activation process limits early HIV replication in a way that may augment current virus-specific ART agents.
For more information, please see Dr. D'Aquila's faculty profile.
Publications
See Dr. D’Aquila's publications in PubMed.
Contact
Phone 312/695-5085

Nicolae Valentin David
Associate Professor, Medicine (Nephrology and Hypertension)
Molecular mechanisms of metabolic bone diseases, especially the regulation and function of FGF23 in normal and abnormal mineral metabolism.

Nicolae Valentin David
Research Area(s): Metabolism and Endocrinology
Dr. David uses a basic science and translational research approach to characterize molecular events that are involved in the expression, post-translational modifications and secretion of the bone hormone FGF23 that is highly elevated in patients with chronic kidney disease (CKD). A major area of his research focuses on investigating a novel mechanism by which inflammatory signals and iron deficiency, common consequences of CKD, regulate FGF23. Our data show that acute inflammation stimulates FGF23 production, but simultaneous increases in FGF23 cleavage maintain normal levels of biologically active protein. However, chronic inflammation and sustained iron deficiency also increase biologically active FGF23, and show that these factors may contribute to elevated FGF23 levels in CKD.
Dr. David’s laboratory is funded by the National Institute of Health, National Institute of Diabetes and the National institute of Digestive and Kidney Diseases (NIDDK).
For more information please see Dr. David's faculty profile.
Publications
See Dr. David's publications on Pub Med.
Contact

Erica E Davis
Professor, Pediatrics, Cell and Developmental Biology
Genetic architecture of rare pediatric disease with emphasis on ciliopathies, undiagnosed rare congenital disorders, and neurodevelopment disorders

Erica E Davis
Research Area(s): Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology
We are focused primarily on the study of pediatric genetic disorders, and our mission is to: a) improve our knowledge of genetic variation that causes these disorders and modulates their severity; b) discover pathomechanisms at the cellular and biochemical level; and c) develop cutting-edge therapeutic modalities that will improve the health and well-being of affected individuals and their families. Our research themes focus on but are not limited to:
- Acceleration of gene discovery in proximal and global pediatric cohorts.
- Understanding the contextual effect of genetic variation to explain pleiotropy, variable expressivity, and epistasis.
- Development and application of experimentally tractable models to delineate underlying pathomechanism.
- Establishing in vitro and in vivo assays for human disease modeling that are suitable for medium and high throughout drug screening.
- Synthesis of clinical investigation and basic experimental biology to advance molecular diagnosis and identify suitable treatment.
For more information, please see Dr. Davis's faculty profile.
Publications
See Dr. Davis's publications in PubMed.
Contact
Phone 312-503-7662

Paul G DeCaen
Associate Professor, Pharmacology
Ion channels in cell biology and disease progression

Paul G DeCaen
Research Area(s): Pharmacology
We study the biophysics, pharmacology and physiology of ion channels. Currently, we are focused on two divergent groups: voltage gated sodium channels (Nav) and Polycystin channels (also called Polycystic Kidney Disease Proteins, PKDs). Aside from these foci, we actively explore novel ion channels found in prokaryotic and eukaryotic cells with the goal of understanding their function in cell physiology.
Current Projects
Voltage Gated Sodium Channels
Navs conduct sodium ions into excitable cells like muscle and neurons, causing the cell membrane to depolarize on the microsecond time scale, a process essential for rapid communication in multicellular organisms. Potentially fatal conditions such as forms of epilepsy and cardiac arrhythmias arise when Navs are mutated.
With our collaborators, we continue to examine key questions:
- How do these transmembrane proteins sense electrical potential and change from nonconductive to conductive states?
- How do these transmembrane proteins select for sodium ions and not allow passage of the other ions present?
- What are the mechanisms of action of clinically relevant drugs (e.g. Valproate and Lamotrigine) and where are their receptor sites?
Polycystin Channels and Primary Cilia
Mutations in PKD1 and PKD2 are associated with Autosomal Dominant Kidney Disease (ADPKD). ADPKD is one of the most common monogenetic diseases in mankind, where progressive cyst formation results in kidney failure. Several members of the polycystins (PKD1, PKD1-L1, PKD2 and PKD2-L1) have been found in the primary cilia from cells of various tissues besides the kidney. The primary cilium is a solitary, small (5-15 mM in length) protuberance from the apical side of polarized cells.
With help from our collaborators, our research is directed to answer key questions:
- How do ADPKD mutations alter PKD2 function? Do some mutations ‘turned on’ while others ‘turn off ’ the PKD2 channel?
- How does PKD1/2 channel dysfunction result in cyst formation? Or conversely, what normal function do they serve for the primary cilium and how do PKDs maintain cell polarity?
- What are the receptor sites within PKD2s that can modulation its ion channel function and are they drug-able?
For lab information and more, see Dr. DeCaen's faculty profile.
Publications
See Dr. DeCaen's publications on PubMed.
Contact
Contact Dr. DeCaen at 312-503-5930.

Alexis Demonbreun
Associate Professor, Pharmacology
Genetic mechanisms of myopathies to drive therapy development.

Alexis Demonbreun
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Pharmacology
The Demonbreun laboratory studies the genetic mechanisms that underlie and modify inherited myopathies. The goal of our research is to use these genetic signals to guide the development of therapeutics for the treatment of cardiac and skeletal muscle injury and disease, with a focus on enhancing membrane repair and reducing fibrosis. Much of this work is oriented towards treating rare genetic diseases that affect heart and skeletal muscle, but the basic pathways targeted by these proteins suggests potential broader clinical utility.
For more information, see Dr. Demonbreun's faculty profile.
Publications
See Dr. Demonbreun's publications on PubMed.
Contact
Contact Dr. Demonbreun

Francesca Elizabeth Duncan
Associate Professor, Obstetrics and Gynecology (Reproductive Science in Medicine)
Mammalian ovarian and gamete biology and reproductive aging

Francesca Elizabeth Duncan
Research Area(s): Developmental and Stem Cell Biology; Reproductive Biology
Aging is associated with cellular and tissue deterioration and is a prime risk factor for chronic
diseases and declining health. The female reproductive system is the first to age in humans, with
a decline in egg quantity and quality beginning at ~35 years of age and menopause ensuing at
~50 years of age. Female reproductive aging has significant health consequences as it results in
endocrine function loss and is a leading cause of infertility, miscarriages, and birth defects.
Although aging hallmarks and mechanisms have been enumerated across multiple organ
systems and species, they have not been investigated in the context of mammalian reproductive
aging.
My research program integrates and builds upon my 18-year history in the field of reproductive
science and medicine to investigate the overarching hypothesis that deterioration of oocyte-intrinsic
cellular pathways together with alterations in the ovarian environment underlie the age-associated
decline in female gamete quantity and quality. Our work is at the interface of
reproductive aging and systemic aging; physiologic and iatrogenic reproductive aging; gamete,
follicle, and ovarian biology; and reproductive science and medicine. Our comprehensive insights
will help us design targeted interventions to potentially slow or counteract reproductive aging,
laying the foundation to simultaneously improve women’s fertile-span and health-span across
generations. In addition, reproductive aging mechanisms may inform those that precipitate
general aging, which occur up to decades later in life. Moreover, the mechanisms involved in
reproductive aging that we are investigating - aneuploidy, protein metabolism dysregulation,
and fibrosis and inflammation – are also central to other conditions such as cancer
pathogenesis. Thus, our research has broad impact and collaborative opportunities across
disciplines, which already include biochemistry, biophysics, toxicology and pharmacology, and
reproductive endocrinology and infertility.
Ultimately our work in reproductive aging will have direct impacts on public health in two ways.
First, reproductive aging affects all women, and menopause and premature aging of the ovary
accelerates aging in general. Such health consequences occur because ovarian hormones such
as estrogen, for example, are critical for cardiovascular, bone, immune, and cognitive functions.
Second, reproductive aging is associated with age-associated infertility, which has significant
societal, clinical, and health ramifications as more women globally are delaying childbearing.
For lab information and more, see Dr. Duncan's faculty profile.
Visit the Duncan Lab website
Publications
See Dr. Duncan's publications on PubMed.
Contact
Email Dr. Duncan
312-503-2172

Igor R Efimov
Professor, Medicine (Cardiology)
Professor, McCormick School of Engineering
Bioengineering approaches to study and treat cardiac arrhythmias

Igor R Efimov
Research Area(s): Cardiovascular and Renal Biology
Cardiac arrhythmias are responsible for 300,000 deaths in the USA. To study and treat arrhythmias, my bioengineering lab pursues bioengineering approaches, including implantable and wearable devices, imaging, ablation, tissue engineering, and machine learning. My trainees, collaborators and I have developed, improved, and applied several experimental methodologies based on optical imaging to study cardiac physiology and pathophysiology. My principal interest is to improve our understanding of the cardiovascular system's development and function and develop novel diagnostics and electrotherapeutics for cardiovascular diseases. I am interested in the mechanisms of arrhythmia and anti-arrhythmic therapy, primarily bioelectronics-based. Our work has resulted in the development of low-energy atrial and ventricular defibrillation by Cardialen, which was acquired in 2022.
During the last decade, in collaboration with Dr. John A. Rogers from Northwestern University, we have developed and tested several novel approaches to organ conformal bioelectronics and medical devices based on flexible and stretchable electronics platforms, including bioresorbable electronics, bioresorbable adhesives, multifunctional catheters, etc. Our current focus is developing novel percutaneous catheter-based diagnostic and therapeutic devices and implantable organ conformal devices, which will provide high-definition diagnostics and therapy for cardiac arrhythmia and failure. Based on this work, we co-founded NuSera Biosystems to commercialize this novel bioelectronic approach.
In 2022, I joined the faculty of Northwestern University as a Professor of Biomedical Engineering and Professor of Medicine (Division of Cardiology) to accelerate our collaborative research within Northwestern’s Querrey Simpson Institute for Bioelectronics and CZ Biohub Chicago.
Some of the currently funded projects:
- Epicardial high-definition electrotherapy of AFib.
- Bioelectronics for Neurocardiology.
- Soft wireless multimodal cardiac implantable devices for long-term investigating heart failure pathogenesis.
- Reagentless Sensor Technologies For Continuous Monitoring of Heart Failure Biomarkers.
- High-Definition Conformal Electronics For VT/VF.
- Optimization of electromechanical monitoring of engineered heart tissues.
- Targeting intracellular calcium leak for ventricular arrhythmia prevention in structural heart disease.
For lab information, publications and more, see Dr. Efimov's faculty profile.
Publications
See Dr. Efimov's publications on PubMed.
Contact
Contact Dr. Efimov.

Lillian J Eichner
Assistant Professor, Biochemistry and Molecular Genetics
Transcriptional dependencies in cancer at the intersection of epigenetics, signaling and metabolism

Lillian J Eichner
Research Area(s): Cancer Biology; Lung Biology
The Eichner lab studies transcriptional dependencies in cancer development, progression, and therapeutic resistance. We endeavor to elucidate causal mechanisms at the intersection of epigenetics, signaling, and metabolism in order to understand tumor biology and harness therapeutically targetable vulnerabilities in cancer.
For lab information, publications and more, see Dr. Eichner's faculty profile and laboratory website.
Publications
See Dr. Eichner's publications on PubMed.
Contact
Contact Dr. Eichner.

Stephanie C Eisenbarth
Professor, Medicine (Allergy and Immunology), Pathology
Cellular and molecular regulation of antibody-mediated disease

Stephanie C Eisenbarth
Research Area(s): Cell Biology; Immunology
The Eisenbarth-Williams lab focuses on defining the cellular and molecular mechanisms that regulate T cell guided antibody responses. The development of antibodies relies on the interactions between three immune cells – dendritic cells (DCs), T cells and B cells. We study how these three cell types communicate to shape different types of antibody responses, some protective in the case of vaccination and some harmful in the case of allergy and alloimmunization. The fundamental principles governing the interaction between these cells are the same across these seemingly disparate responses, yet specialization in the subset of each cell type, the specific niche for the interaction and the cellular signals exchanged between the cells dictates what type of antibody response is generated. More recently, our work has expanded to include the mucosal epithelium, which influences the function of these three immune cell types. By utilizing human samples to guide our studies and mouse models to test new mechanistic paradigms, we have identified novel and unexpected cell subsets and functions. Ultimately, this knowledge can be harnessed to induce protective immune responses and subvert pathogenic ones.
For lab information, publications and more, see Dr. Eisenbarth's faculty profile.
Publications
See Dr. Eisenbarth's publications on PubMed.
Contact
Contact Dr. Eisenbarth.

Elizabeth A Eklund
Professor, Medicine (Hematology and Oncology)
Myeloid leukemia and approaches to chemotherapy resistant disease.

Elizabeth A Eklund
Research Area(s): Genetics, Epigenetics and Genomics; Pharmacology
Dr. Eklund’s laboratory studies are focused on understanding the molecular events that lead to development of myeloid leukemias (acute myeloid leukemia and chronic myeloid leukemia) and to the evolution of drug resistance in these diseases. The goal is to identify potential molecular therapeutic targets that would delay or prevent drug resistance and relapse in AML and CML. In related projects, the laboratory is investigating Fanconi Anemia, a genetic disease with defective DNA repair. Patients with Fanconi Anemia frequently develop leukemia and provide a model for understanding the role of DNA repair in leukemogenesis.
Publications
View lab publications via PubMed.
For more information, visit the faculty profile page of Elizabeth Eklund, MD.
Contact Us
Contact Dr. Eklund at 312-503-3208 or the Eklund Lab at 312-503-3208.

Deyu Fang
Professor, Pathology (Experimental Pathology)
Molecular networks in the regulation of immune response and autoimmunity

Deyu Fang
Research Area(s): Immunology
Our research goal is to identify the therapeutic molecular targets for the treatment of autoimmune diseases particularly of rheumatoid arthritis (RA) and type 1 diabetes (T1D).
In our laboratory, we use genetic, proteomic, molecular biology and immunological approaches to dissect the molecular networks underlying the regulation of immune response and autoimmunity. Several specific genes that are critical for immune regulation and autoimmune diseases have been identified in our laboratory. Small molecules that modulate the functions of these newly identified genes can potentially be used to treat type 1 diabetes and rheumatoid arthritis.
The current ongoing research projects, in my laboratory are:
1. Sirt1, a type-iii histone deacetylase, is required for immune tolerance.
2. The ubiquitin E3 ligase Synoviolin, is a therapeutic target for RA.
3. The tyrosine kinase c-Abl in T-cell differentiation and allergic lung inflammation.
4. The roles of RoxP3 in regulatory T cells.
5. Ubiquitination in aging and autoimmunity.
6. Novel microRNAs in immune tolerance and autoimmunity.
For more information, visit Dr. Deyu Fang's faculty profile or laboratory site.
Publications
See Dr. Fang's publications in PubMed.
Contact

Amani A Fawzi
Professor, Ophthalmology
Molecular mechanisms of ischemic retinopathies and retinal fibrosis- the neurovascular perspective

Amani A Fawzi
Research Area(s): Cell Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Neurobiology
My research lab focuses on translational, basic science projects that aim to identify the critical driving factors in the vascular pathology in diabetic retinopathy (DR). We have a special focus the pathogenesis of fibrovascular transition in DR, which is a devastating clinical stage. Very little is currently known about the drivers of fibrovascular pathology in the ischemic retina, which is a vision threatening end stage of many ischemic retinopathies.
We are taking a systems biology approach to this problem. While most researchers have focused on one isolated compartment, either the retinal neurons, glia or vessels, our approach takes all these players into account in order to construct a wholistic view of the ischemic retina and diabetic retinopathy. Using single- and bulk RNA seq, we have collected a large dataset from mouse and human diabetic tissue. We are looking for candidates who are interested in studying this disease in depth and who have an interest in bio-informatics, RNA-Seq and neurovascular interactions.
For more information, visit the faculty profile for Amani A. Fawzi, MD.
Publications
See Dr. Fawzi's publications on PubMed.
Contact

Adriana B Ferreira
Associate Professor, Cell and Developmental Biology
Mechanisms underlying neurite degeneration and synapse loss in Alzheimer’s disease and related neurodegenerative disorders

Adriana B Ferreira
Research Area(s): Cell Biology; Neurobiology
Our work focuses on the mechanisms underlying neurite degeneration and synapse loss in Alzheimer’s disease and related neurodegenerative disorders. We are interested in the relationship between beta-amyloid deposition and the progressive formation of dystrophic neurites and cell death in hippocampal neurons.
Recently, we have determined that the microtubule associated protein tau plays an essential role in beta-amyloid-induced neurite degeneration. These results constitute the first direct evidence of a mechanistic link between beta-amyloid deposition and tau in central neurons. Furthermore, our results indicated that beta-amyloid induces calpain-mediated tau cleavage leading to the generation of a 17 kDa neurotoxic fragment in hippocampal neurons both in culture model systems and in AD human brain samples. Currently, we are analyzing the mechanisms by which this tau fragment mediates beta-amyloid-induced neurite degeneration. These studies are being performed by means of a variety of cell and molecular biology techniques.
For lab information and more, see Dr. Ferreira's faculty profile and lab website.
Publications
See Dr. Ferreira's publications on PubMed.
Contact
Contact Dr. Ferreira at 312-503-8250.

Daniel R Foltz
Professor, Biochemistry and Molecular Genetics
Epigenetic control of centromere assembly and chromosome segregation

Daniel R Foltz
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics
My research program is focused on the important basic question of how chromosomes are segregated during cell division to ensure the complete and accurate inheritance of the genome. Chromosome instability is a hallmark of cancer and can drive tumorigenesis. Therefore, how centromere specification is controlled is a basic biological question with great therapeutic potential. Centromeres are specified by the incorporation of a histone variant CENP-A in a centromere specific nucleosome. The stable inheritance of this locus is controlled by an epigenetic pathway and does not depend on the underlying DNA sequence. My research program is using a combination of cell biology, biochemical purification and in vitro reconstitution of centromeric chromatin to discover the mechanisms of epigenetic inheritance and CENP-A function during mitosis. A key to understanding the epigenetic inheritance of centromeres is determining the process by which new CENP-A nucleosomes are deposited. Our lab is studying how activity of the CENP-A chromatin assembly factor HJURP is coupled to existing centromeres. Non-coding RNAs, as well as chromatin modifying enzymes have been implicated in the process and we are exploring how these factors contribute to specific assembly of the CENP-A nucleosomes. We have identified novel post translational modifications of the CENP-A amino-terminus and we are working to determine how these modifications contribute to genomic stability and accurate chromosome segregation. Our immediate goal is to determine the mechanism of epigenetic centromere inheritance, with a long-term goal of delineating the role of this process in tumorigenesis so as to translate our basic understanding of the enzymes and proteins involved in this process into therapeutic approaches for genomic instability in cancer.
For lab information and more, see Dr. Foltz's faculty profile.
Publications
See Dr. Foltz's publications on PubMed.
Contact
Contact Dr. Foltz at 312-503-5684.

Slim Fourati
Assistant Professor, Medicine (Allergy and Immunology)
Integrating High-Throughput Data for Predictive Modeling of Immune Responses

Slim Fourati
Research Area(s): Computational Biology and Bioinformatics; Immunology; Microbiology; Pharmacology
My research focuses on the interplay between host and pathogens, focusing on the use of
bioinformatic analysis of high-throughput data (like RNA-Sequencing, Flow cytometry) to identify
biological mechanisms associated with the immune response to pathogens, the use of machine
learning to build models able to predict severity of the symptoms following infections or
response to prophylactic therapies against those same pathogens and the use of in silico
modeling to identify new therapeutic intervention.
For lab information and more, see Dr. Fourati's faculty profile
Publications
See Dr. Fourati's publications on PubMed.
Contact
Contact Dr. Fourati

Colin K Franz
Associate Professor, Physical Medicine and Rehabilitation, Medicine (Pulmonary and Critical Care), Neurology - Ken and Ruth Davee Department, Physical Therapy and Human Movement Sciences
Translational neuromuscular medicine from human neural organoids to improving clinical outcomes

Colin K Franz
Research Area(s): Cell Biology; Developmental and Stem Cell Biology; Neurobiology
Dr. Colin Franz, MD, PhD, is a physician-scientist with board certifications in neuromuscular medicine, electrodiagnostic medicine, and physical medicine and rehabilitation. He serves as a staff physician at the Lois Insolia ALS Clinic at Northwestern Memorial Hospital and as the director of both the Regenerative Neurorehabilitation Laboratory (research) and the Electrodiagnostic Laboratory (clinical) at Shirley Ryan AbilityLab. Dr. Franz's clinical interests encompass the management of complex peripheral nerve injuries, amyotrophic lateral sclerosis (ALS), and diaphragm muscle dysfunction, alongside clinical neurophysiology and innovative applications of neuromuscular ultrasound. His research team employs a technology-driven approach to personalize rehabilitation medicine, inspired by the real-life challenges faced by his patients. Dr. Franz's work includes pioneering studies, from developing next-generation nerve implants from advanced materials for targeted therapeutics and nerve regeneration, to creating human neural organoids from patient-derived pluripotent stem cells to investigate motor neuron degeneration post-spinal cord injury. These initiatives aim to understand how patient-specific factors influence neurotrauma outcomes and the potential for nerve regeneration. Dr. Franz's research is robustly supported by prestigious external institutions, including the National Institutes of Health (NIH), the American Neuromuscular Foundation, the National Science Foundation (NSF), and the Belle Carnell Regenerative Neurorehabilitation Fund.
For lab information and more, see Dr. Franz's faculty profile.
Publications
See Dr. Franz's publications on PubMed.
Contact
Contact Dr. Franz

William E Funk
Associate Professor, Preventive Medicine (Epidemiology)
Biomarker discovery and biomarker applications to epidemiological research

William E Funk
Research Area(s): Cancer Biology
Recent findings suggest that more than 70% of the risk for developing chronic diseases and cancers are due to environmental factors, which include interactions between our environment and our genes. However, the role of the environment in disease etiology remains largely unknown. To address this need, research in my laboratory is focused on biomarker discovery and targeted methods that utilize simple and non-invasive blood collection strategies to extend biomarker applications to epidemiological research.
A major focus of my research is on an important class of chemical toxicants, reactive electrophiles. Because electrophiles are reactive, adducts of abundant blood proteins are used as biomarkers of exposure. Since stable adducts persist in the blood for the life span of the protein, adducts reflect an integration of exposure over weeks to months. We are applying an “adductomics” approach to a single nucleophilic ‘hot spot” on human serum albumin (HSA), HSA-Cys34, to generate adduct maps as molecular fingerprints of exposure. These adduct maps can be compared across groups to pinpoint candidate biomarkers. We are currently applying this strategy to investigate unknown risk factors associated with several cancers, congenital disorders and chronic diseases, including ovarian cancer, multiple myeloma, cleft lip and cleft palate, spina bifida and cardiovascular disease.
A second line of research in my laboratory focuses on the logistics of measuring biomarkers to assess environmental exposures in population-based research. An obstacle to the measurement of biomarkers is the requirement for venipuncture to obtain samples, which is expensive, invasive and needs to be performed by a trained phlebotomist near a laboratory where the blood can be immediately processed. In contrast, dried blood spots (DBS)—drops of whole blood collected on filter paper—are a simple and non-invasive alternative that can be collected in non-clinical settings. Current DBS projects in our laboratory are focused on prenatal, early childhood and adult exposures to toxic metals, including lead, arsenic, mercury and cadmium. We have several ongoing collaborative projects applying DBS to assess heavy metal exposures in newborns using state-archived DBS (retrospective studies) and in pediatric populations and young children (prospective studies) using a newly developed blood collection device (Funk & McDade, Patent Pending).
Future studies are planned to combine both lines of my research and apply adductomics to DBS samples. In collaboration with the CDC’s Centers for Birth Defects Research and Prevention, we will be obtaining newborn DBS from State Public Health Laboratories to explore unknown prenatal risk factors for a variety of birth defects.
For more information visit the faculty profile of William Funk, PhD.
Publications
See Dr. Funk's publications in PubMed.
Contact

Ruli Gao
Assistant Professor, Biochemistry and Molecular Genetics
Tumor evolutionary models and immunological reprogramming with new computational tools and long read single-cell sequencing technology

Ruli Gao
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics
The Gao laboratory at Northwestern University studies genetic evolutionary models and cell fate transition trajectories during cancer progression and therapeutic responses. The laboratory has a focus on developing novel long-read single cell sequencing technologies and new computational tools. Previous studies include the development of the punctuated copy number evolution model ( Nature Genetics, 2016), the bi-mode chemoresistance evolution model (Cell, 2018), the high throughput single nuclei RNA sequencing method (Nature Communications, 2017), the computational algorithm, COPYKAT for calculating single cell copy number clonality from scRNA-seq data (Nature Biotechnology, 2021), the long-read single cell sequencing technology and computational tool, scNanoGPS (Nature Communications, 2023) and the intra-tumor anaplastic transformation spectrum (JCI, 2023). The ongoing projects are centered around integrating same-cell multi-omics to co-project tumor cell genetic evolutionary lineages, cell fate transition trajectories and adaptive immunological remodeling during cancer metastasis and recurrence. Additionally, the laboratory is expanding the applications of these novel methodologies onto diverse human disease models, such as heart aging, heart failure and transplantation rejection.
For more information, please see Dr. Gao's faculty profile or the Gao lab website.
Publications
See Dr. Gao's publications in PubMed.
Contact
Contact Dr. Gao at 312-503-3796.

Jaime Garcia-Anoveros
Professor, Anesthesiology, Neuroscience, Neurology - Ken and Ruth Davee Department
Development, function, dysfunction and degeneration of sensory receptor cells and neurons

Jaime Garcia-Anoveros
Research Area(s): Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology
We study the development of the cochlea and its innervation because it is a model organ in which to elucidate fundamental principles of development and neuroscience. Our research aims at overcoming the obstacles that lay in the way towards therapeutic regeneration and restoration of lost hearing. Current projects address how inner and outer sensory hair cells form, how they assemble with specialized supporting cells into the cellular mosaic that is the organ of Corti, and how they become innervated so as to communicate with the brain.
Development of inner and outer hair cells: The two mechanosensory cells in the cochlea, inner (IHC) and outer (OHC) hair cells, play complementary roles in hearing. Cochlear hair cells are produced exclusively during an early developmental period, and hence their death throughout life, whether caused by age, noise or ototoxic drugs, result in irreversible deafness. Attempts at generating new hair cells from cochlear supporting cells are under way, but we need to know how to specifically generate IHCs and OHCs, each in their appropriate location. We found that this involves a morphogen, a transcription factor that renders OHCs insensitive to it, and a master regulator that induces IHC formation and maintenance (Wiwatpanit et al., 2018, Nature; García-Añoveros et al., 2022, Nature). Having identified the major regulators, we are now able to elucidate how they execute their molecular functions and to use this information to drive the therapeutic generation of new hair cells into specifically forming IHCs or OHCs.
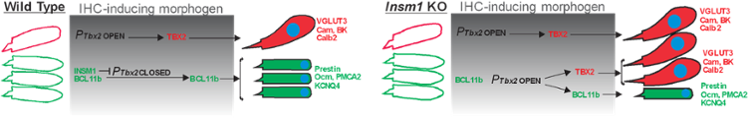
Figure 1: Summary of the molecular mechanisms for the generation of IHCs vs OHCs.
Intercellular arrangement of hair and specialized supporting cells. We are also exploring how the specialized supporting cells for IHCs and OHC are generated and assemble with them into the near crystalline mosaic that is the organ of Corti. This basic information will be critical for any regenerative attempt to restore hearing; it will also reveal fundamental mechanisms for how multiple cell types assemble to make an organ.
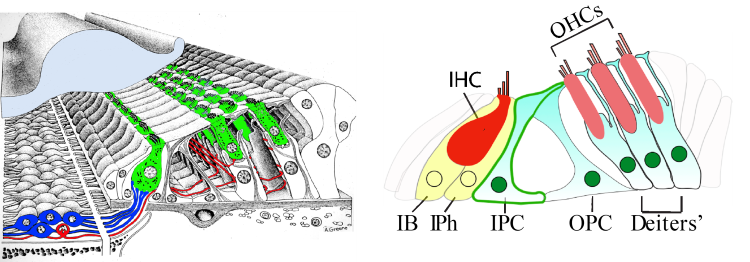
Figure 2: Mosaic intercellular arrangement of the organ of Corti and indication of the specialized supporting cell types paired with, and separating, IHCs and OHCs.
Assembly of cochlear neural circuits. The cochlea communicates with the brain via an iterative but relatively simple circuit composed of a distinct pair of afferent plus efferent neurons for IHCs, and another for OHCs. Taking advantage of our ability to interconvert IHCs into OHCs and vice versa, we begun to discover general steps for the formation of this circuit (Webber et al., 2021, Science Advances). We expect to elucidate first the general principles, and then the molecular mechanisms, guiding the development of cochlear innervation, essential for hearing and paradigmatic for neuroscience.
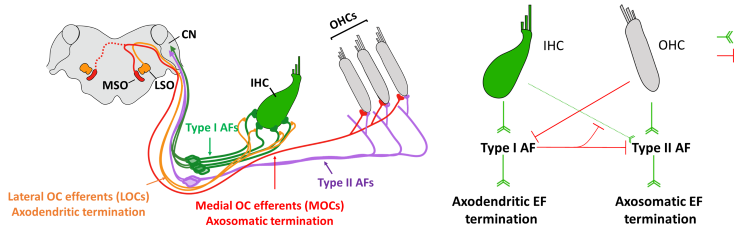
Figure 3: Innervation of the cochlea and the sequence of signals (attractive in green and repulsive in red) by which it is achieved.
For more information, view the faculty profile of Jaime García-Añoveros, PhD or visit the Añoveros & Duggan lab site.
Publications
See Dr. García-Añoveros' publications on PubMed.

David M Gate
Assistant Professor, Neurology (Behavioral Neurology)
Intersection of the immune system and neurodegenerative disease

David M Gate
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Immunology; Neurobiology
The Gate lab in Northwestern Neurology works at the interface of the immune system and neurodegenerative disease. The lab is focused on employing human genomics approaches to uncover novel biomarkers and therapeutic targets for neurodegeneration. Chief among our strategies is single cell RNA sequencing (scRNAseq) to identify transcriptional changes in human specimens. We also employ spatial transcriptomics, immunohistochemistry and cytometry approaches to validate genomic changes observed by scRNAseq. The Gate lab is focused primarily on neurodegenerative diseases of aging, including, but not limited to: Alzheimer’s disease, Parkinson’s disease and Amyotropic lateral sclerosis.
For lab information and more, see Dr. Gate's faculty profile.
Publications
See Dr. Gate's publications on PubMed.
Contact
Email David Gate, PhD

Vladimir I Gelfand
Professor, Cell and Developmental Biology
Molecular motors and organelle movement

Vladimir I Gelfand
Research Area(s): Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology; Reproductive Biology
One of the remarkable features of eukaryotic cells is their ability for rapid transport of intracellular organelles in the cytoplasm. Examples of such transport include segregation of chromosomes during cell division and the transport of organelles in neurons from the cell body into axons and dendrites. Movements of organelles are powered by molecular motors. Microtubule motors(kinesins and dyneins) move along microtubules and myosins move along microfilaments.
We use two cellular models to discover how multiple motors on the surface of the same cargo work together in organelle movement and how these motors are attached to the surface of organelles and what regulates their activity. One model is cultured pigment cells (melanophores). These cells activate movement of pigment organelles in response to hormone-modulated changes of cAMP concentration. The movement of pigment organelles is powered by three different motors (two microtubule motors of different polarity and a myosin) and this system is very convenient for analysis of motor regulation. A second model is cultured Drosophila cells that we use to individual components of transport machinery by using RNAi. In our work, we employ techniques of cell and molecular biology and computer-assisted microscopy of living cells and purified organelles as well as high-resolution and high-sensitivity biophysical methods.
For lab information and more, see Dr. Gelfand's faculty profile and lab website.
Publications
See Dr. Gelfand's publications on PubMed.
Contact
Contact Dr. Gelfand at 312-503-0530.

Alfred L. George, Jr.
Professor, Pharmacology
Structure, function, pharmacology and molecular genetics of ion channels and channelopathies

Alfred L. George, Jr.
Research Area(s): Cardiovascular and Renal Biology; Genetics, Epigenetics and Genomics; Neurobiology; Pharmacology

Research Description
Ion channels are ubiquitous membrane proteins that serve a variety of important physiological functions, provide targets for many types of pharmacological agents and are encoded by genes that can be the basis for inherited diseases affecting the heart, skeletal muscle and nervous system.
Dr. George's research program is focused on the structure, function, pharmacology and molecular genetics of ion channels. He is an internationally recognized leader in the field of channelopathies based on his important discoveries on inherited muscle disorders (periodic paralysis, myotonia), inherited cardiac arrhythmias (congenital long-QT syndrome) and genetic epilepsies. Dr. George’s laboratory was first to determine the functional consequences of a human cardiac sodium channel mutation associated with an inherited cardiac arrhythmia. His group has elucidated the functional and molecular consequences of several brain sodium channel mutations that cause various familial epilepsies and an inherited form of migraine. These finding have motivated pharmacological studies designed to find compounds that suppress aberrant functional behaviors caused by mutations.
Recent Findings
- Discovery of novel, de novo mutations in human calmodulin genes responsible for early onset, life threatening cardiac arrhythmias in infants and elucidation of the biochemical and physiological consequences of the mutations.
- Demonstration that a novel sodium channel blocker capable of preferential inhibition of persistent sodium current has potent antiepileptic effects.
- Elucidation of the biophysical mechanism responsible for G-protein activation of a human voltage-gated sodium channel (NaV1.9) involved in pain perception.
Current Projects
- Investigating the functional and physiological consequences of human voltage-gated sodium channel mutations responsible for either congenital cardiac arrhythmias or epilepsy.
- Evaluating the efficacy and pharmacology of novel sodium channel blockers in mouse models of human genetic epilepsies.
- Implementing high throughput technologies for studying genetic variability in drug metabolism.
- Implementing automated electrophysiology as a screening platform for ion channels.
For lab information and more, see Dr. George’s faculty profile.
Publications
See Dr. George's publications on PubMed.
Contact
Contact Dr. George at 312-503-4892.

Richard C Gershon
Professor, Medical Social Sciences (Outcome and Measurement Science)
Health care outcome assessments

Richard C Gershon
Research Area(s): Computational Biology and Bioinformatics
Dr. Gershon is a leading expert in the application of Item Response Theory (IRT) in individualized and large scale assessments. He has developed item banks and Computerized Adaptive Testing (CAT) for educational, clinical, and health applications - including cognitive, emotional, and motor applications. He is currently principal investigator on these projects with the NIH: NIH Toolbox for the Assessment of Neurological Function and Behavior, the NIH Roadmap Patient – Reporting Outcomes Measurement Information System (PROMIS) Technical Center, the National Institutes on Aging Genetic Norming project, and the National Children's Study: Vanguard Study(South ROC). He is also co-investigator and measurement development expert on numerous smaller projects including the NINDS sponsored project “Quality of Life Outcomes in Neurological Disorders” (Neuro-QOL), and the cancer-specific supplement to PROMIS.
For more information visit the faculty profile of Rich Gershon, PhD.
Publications
See Dr. Gershon's publications in PubMed.
Contact

Lucy A Godley
Professor, Medicine (Hematology and Oncology)
Germline cancer risk and epigenetic control of cell differentiation

Lucy A Godley
Research Area(s): Cancer Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Immunology
Dr. Godley received scientific training in the Marchesi (Yale), Wiley (Harvard), and Varmus (UCSF/NIH) laboratories and medical training at Northwestern. After training in Internal Medicine and Hematology-Oncology at The University of Chicago, she held a long-time faculty position at the University of Chicago and recently relocated to Northwestern University where she is the Jeffrey and Marianne Silver Family Professor of Oncology; Director of the Silver Family Blood Cancer Institute and Clinical Director of Cancer Genetics for the Robert H. Lurie Comprehensive Cancer Center and Division of Hematology/Oncology. Dr. Godley also runs an independent research laboratory focused on understanding the molecular pathways that drive hematopoietic malignancies, especially how germline predisposition alleles contribute to individual and family risk. Dr. Godley has contributed to the recognition of germline DDX41, ETV6, and CSF3R variants as risk factors to developing hematopoietic malignancies. She is also studying how deleterious germline RUNX1, CHEK2, and BRCA1/2 variants drive these cancers, especially considering how the development of clonal hematopoiesis and inflammatory pathways contribute to tumorigenesis. Dr. Godley has shown that germline contribution to hematopoietic malignancies occurs throughout the entire age range of life and is more common than previously anticipated, which has important implications for the allogeneic hematopoietic stem cell donor pool. Along with David Wu (Univ of Washington), Dr. Godley co-chairs the Myeloid Malignancy Variant Curation Expert Panel, which has provided variant curation rules for RUNX1 that are now in use throughout the world and is currently developing similar rules for GATA2 and DDX41. Dr. Godley’s group also studies how covalently modified cytosines in DNA control cellular differentiation, specifically how hypoxia alters epigenetic modifications with hematopoietic stem and progenitor cells, especially during erythropoiesis. Dr. Godley practices bench to bedside medicine that advances patient care through a deep appreciation of the latest scientific findings.
To learn more, please visit Dr. Godley's faculty profile
Publications
See the Dr. Godley's publications on the PubMed.
Contact

Robert Goldman
Professor Emeritus, Cell and Developmental Biology
Intermediate filament (IF) system in fibroblasts, epithelial cells and nerve cells

Robert Goldman
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Cell Biology; Neurobiology
We focus on the structure and function of cytoskeletal systems, particularly the intermediate filament (IF) system in fibroblasts, epithelial cells and nerve cells. IFs are composed of large families of proteins that vary in composition from one cell type to another, even among cells in the same tissue. Using a variety of techniques, we have demonstrated that IFs form elaborate networks that course throughout the cytoplasm and establish connections with both the nuclear and cell surfaces.
At the nuclear surface, they are linked either directly or indirectly with the nuclear lamins, which are chromatin-associated IF protein family members. At the level of the plasma membrane, IFs are involved as cytoskeletal linkages to the focal adhesion of fibroblasts and the desmosomes and hemidesmosomes of epithelial cells. Throughout the cytoplasm, we have shown that IFs are associated with the other cytoskeletal elements, such as microtubules and microfilaments.
Our approach to studying the IF system involves biochemical, morphological, immunological, cell physiological and molecular techniques. Our hypothesis is that the IF system forms a continuous network linking the nuclear and cell surfaces, functioning in such diverse activities as the establishment and maintenance of cell shape, organelle movements within the cytoplasm, nuclear positioning, nuclear-cytoplasmic interactions and signal transduction.
Since many human diseases have been linked to changes in cytoskeletal IF systems, we are also developing models to study the mechanisms involved in IF alterations in various diseases. One example is amyotrophic lateral sclerosis (ALS or Lou Gehrig's disease) in which we have been able to induce neurofibrillary tangles to form in single cultured nerve cells. These tangles are similar to those found in ALS neurons. Therefore, we are able to study the effects of neurofilament tangle formation in single cells. During the summer, researchers from this laboratory also conduct studies on the mechanisms of chromatin/nuclear envelope interactions in eggs of the surf clam at the Marine Biological Laboratory in Woods Hole.
For lab information and more, see Dr. Goldman's faculty profile and lab website.
Publications
See Dr. Goldman's publications on PubMed.
Contact
Contact Dr. Goldman at 312-503-4215.

Jeffery A Goldstein
Associate Professor, Pathology (Perinatal Pathology), Pathology (Autopsy)
Placental diagnosis and deep phenotyping using machine learning and artificial intelligence

Jeffery A Goldstein
Research Area(s): Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Reproductive Biology
For lab information and more, see Jeffrey Goldstein, MD,PhD, faculty profile.
Publications
See Dr. Goldstein's publications
Contact
Email Dr. Goldstein

Cara J Gottardi
Professor, Medicine (Pulmonary and Critical Care)
How cells adhere to each other and how this adhesion is regulated and controls gene expression in heath and disease

Cara J Gottardi
Research Area(s): Cancer Biology; Cell Biology; Lung Biology
The ability of individual cells to adhere and coalesce into distinct tissues is a major feature of multicellular organisms. Research in my laboratory centers on a protein complex that projects from the cell surface and forms a structural “Velcro” that holds cells to one another. This complex is comprised of a transmembrane “cadherin” component that mediates Ca++-dependent homophilic recognition and a number of associated “catenins” that link cadherins to the underlying cytoskeleton. A major focus in our lab is to understand how these catenins direct static versus fluid adhesive states at the plasma membrane, as well as gene expression and differentiation in the nucleus. These basic questions are shedding new light on how dysregulation of the cadherin/catenin adhesion system drives pathologies such as asthma, fibrosis and cancer.
For more information, please see Dr. Gottardi's faculty profile or the Gottardi lab website.
Publications
See Dr. Gottardi's publications in PubMed.
Contact

Eva Henriette Gottwein
Associate Professor, Microbiology-Immunology
Molecular biology of Kaposi's Sarcoma-associated herpesvirus and its associated cancers

Eva Henriette Gottwein
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics; Microbiology
Viruses commonly modify their cellular environment to optimize viral replication and persistence. Much has been learned about the intervention of viral proteins with cellular pathways. More recently, it has become clear that herpesviruses also encode large numbers of microRNAs (miRNAs). The modest amount of space miRNA precursors occupy in the viral genome, their lack of immunogenicity and their potential as regulators of gene expression make miRNAs ideal candidates for viral effectors.
The lab’s research focuses on identifying targets and functions of miRNAs encoded by the human herpesvirus Kaposi’s sarcoma-associated herpesvirus (KSHV). KSHV causes cancer in immuno-compromised individuals. The clinically most relevant KSHV-induced disease is Kaposi’s sarcoma (KS), a complex tumor driven by KSHV-infected endothelial cells. Due to the AIDS epidemic, KS has become the most common cancer in parts of Africa. KSHV also infects B lymphocytes and can consequently cause B cell lymphomas, including primary effusion lymphoma (PEL). KSHV constitutively expresses viral miRNAs from 12 precursors, suggesting a role of these miRNAs in viral replication and pathogenesis.
My lab is currently pursuing the comprehensive identification of mRNA targets of these miRNAs in primary effusion lymphoma cell lines and KSHV-infected endothelial cells. Our data suggest that, together, the KSHV miRNAs directly target hundreds of cellular mRNAs encoding proteins with roles in several different biological pathways. Our goal is to use this knowledge to characterize the most important functions of the KSHV miRNAs.
For lab information and more, see Dr. Gottwein's faculty profile and lab website.
Publications
See Dr. Gottwein's publications on PubMed.
Contact
Contact Dr. Gottwein at 312-503-3075 or the lab at 312-503-3076.

Yogesh Goyal
Assistant Professor, Cell and Developmental Biology
Assistant Professor, McCormick School of Engineering
Single-cell biology of development and disease

Yogesh Goyal
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics
We are interested in understanding the control principles governing biological processes at various scales, from tissue morphogenesis in development to cell-fate decisions in single cells. In particular, individual cells within a genetically identical population constantly undergo fluctuations in their molecular state which may enable them to adopt new fates in response to stimuli. As opposed to hardwired responses encoded in the genome, this dynamic ability of cells to react, either alone or in concert with each other, to external and internal cues is broadly referred to as “plasticity”. Leveraging our unique interdisciplinary approach, we combine novel frameworks, both experimental (e.g. single-cell profiling, spatiotemporally-resolved lineage tracing, multiplexed single-molecule RNA imaging, optogenetics) and computational (stochastic modeling, statistical analysis of large datasets), to track and control state-to-fate mappings in the context of animal development and cancer progression. Working across multiple mammalian model systems (cell lines, developmental and tumor organoids, mouse models), we will ask how diverse non-genetic variabilities drive fate choices and self-organization in development and disease.
To learn more, please visit Dr. Goyal's faculty profile or lab website.
Publications
See the Goyal Lab’s publications on the Goyal Lab Website.
Contact

Kathleen J Green
Professor, Pathology (Experimental Pathology), Dermatology
Cell-to-cell adhesion molecules' integration of mechanical and signaling functions in skin and heart differentiation, disease and cancer

Kathleen J Green
Research Area(s): Cancer Biology; Cell Biology
Dr. Green's research program focuses on how cell-cell adhesion molecules and their associated proteins integrate mechanical and chemical signaling pathways to contribute to the development and maintenance of multicellular tissues. In particular they are investigating how specialized intercellular junctions called desmosomes are assembled and function in ways that transcend their classic textbook definition as spot welds. The lab has shown that desmosomal cadherins help control the balance of proliferation and differentiation and even regulate the production of cytokines that participate in paracrine signaling. Loss of this “brake” results in increased allergic and inflammatory pathways that underlie pathogenesis in inherited disease and possibly cancer, including melanoma. Desmosomes also integrate the functions of other intercellular junctions including gap junctions and interfering mutations can cause lethal heart arrhythmias.
The lab uses a multi-faceted approach, including but not limited to collaborative atomic structure determinations, molecular genetics, live cell imaging, human tissue engineering and gene targeting approaches. Dr. Green serves as Associate Director for Basic Sciences in the R.H. Lurie Comprehensive Cancer Center.
For more information, please visit the Green Lab website and the faculty profile of Kathleen J Green, PhD.
Publications
See Dr. Green's publications in PubMed.
Contact

Richard Green
Professor, Medicine (Gastroenterology and Hepatology)
Genetics and molecular biology of cholestatic liver diseases and fatty liver disorders and the role of ER Stress and the Unfolded Protein Response (UPR) in the pathogenesis of these diseases

Richard Green
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology
Dr. Green’s laboratory investigates the mechanisms of cholestatic liver injury and the molecular regulation of hepatocellular transport. Current studies are investigating the role of the UPR in the pathogenesis and regulation of hepatic organic anion transport and other liver-specific metabolic functions. We employ genetically modified mice and other in vivo and in vitro models of bile salt liver injury in order to better define the relevant pathways of liver injury and repair; and to identify proteins and genes in these pathways that may serve as therapeutic targets for cholestatic liver disorders.
The laboratory also investigates the mechanisms of liver injury in fatty liver disorders and the molecular regulation of hepatic metabolic pathways. The current focus of these studies includes investigations on the role of the UPR in the pathogenesis of non-alcoholic steatohepatitis and progressive fatty liver disease. We employ several genetically modified mice and other in vivo and in vitro models of fatty liver injury and lipotoxicity. Additional studies include the application of high-throughput techniques and murine Quantitative Trait Locus (QTL) analysis in order to identify novel regulators of the UPR in these disease models.
Publications
See Dr. Green's publications in PubMed.
For more information, please see Dr. Green's faculty profile.
Contact
Contact Dr. Green at 312-503-1812 or the Green Lab at 312-503-0089

Alicia D Guemez Gamboa
Assistant Professor, Neuroscience
Novel molecular bases of cellular recognition that govern neuronal circuit assembly during human development and disease

Alicia D Guemez Gamboa
Research Area(s): Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology
Developing neurons integrate into functional circuits through a series of cell recognition events, which include neuronal sorting, axon and dendrite patterning, synaptic selection, among others. Our research focuses on cell-surface recognition molecules that mediate interactions between neurons to discriminate and select appropriate targets in the developing brain. Additionally, we seek to uncover novel mechanisms of neural recognition that lead to brain connectivity defects in humans. To explore the broader roles for cell recognition molecules and their pivotal function in neural circuit development, our lab takes advantage of a battery of modern laboratory techniques. These approaches include animal and stem cell disease modeling, as well as next-generation sequencing and CRISPR/Cas9 gene editing. Identifying fundamental principles of cellular recognition in wiring circuits contributes to our understanding of neurological disorders and how neuronal dysfunction arises from aberrations during development of the human brain.
For lab information and more, see Dr. Guemez-Gamboa's faculty profile and lab website.
Publications
See Dr. Guemez-Gamboa's publications on PubMed.
Contact
Contact Dr. Guemez-Gamboa at 312-503-0752.

SeungHye Han
Assistant Professor, Medicine (Pulmonary and Critical Care)
Mitochondrial metabolism and lung stem cells in lung injury and repair

SeungHye Han
Research Area(s): Cell Biology; Developmental and Stem Cell Biology; Lung Biology
The Han lab studies how mitochondrial metabolism regulates lung stem/progenitor cells by focusing on the different functions of each mitochondrial electron transport chain complex. We are particularly interested in how lung epithelial cells repair after injury and what factors lead to aberrant repair resulting fibrosis. Mitochondrial dysfunction is commonly observed in patients with severe pneumonia and/or pulmonary fibrosis. Using elaborately designed genetic knock-out and knock-in mouse systems, we are testing whether mitochondrial electron transport chain complexes are necessary for proper lung epithelial stem/progenitor cell differentiation. We use viral pneumonia as a model of lung injury and repair in animals. Findings from these studies can be directly applied to the patients hospitalized for pneumonia, influenza, and SARS-CoV-2 infection in our hospital.
For more information, see Dr. Han's faculty profile.
Publications
See Dr. Han's publications on PubMed
Contact

Alan R Hauser
Professor, Microbiology-Immunology, Medicine (Infectious Diseases)
Pathogenesis of Hospital-Acquired Multidrug-Resistant Bacteria

Alan R Hauser
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Lung Biology; Microbiology
We are using genomic approaches to examine why some bacterial strains within a species are more virulent or transmissible than others. These investigations have been applied to multidrug-resistant bacteria such as Pseudomonas aeruginosa and Klebsiella pneumoniae. To date, studies have led to the identification of several novel virulence factors such as CdiA, which is a contact-dependent inhibition protein that represents a new family of virulence factors. Other studies have identified “high-risk” strains of multidrug resistant bacteria that have spread locally at our hospital or globally. We are expanding these studies to identify additional novel virulence, colonization, and antibiotic resistance factors that will then be characterized using molecular techniques, microscopy, tissue culture and mouse models to explore their mechanisms.
In a second project, we investigate the role of the type III secretion system of P. aeruginosa in pathogenesis. Type III secretion pathways form needle-like apparatuses that inject toxins directly into the cytoplasm of host cells. They are used by many gram-negative bacteria to injure or kill human cells and thus cause disease. We focus on the interaction of the secreted toxins with the host. Current projects involve the use of molecular and biochemical techniques to identify novel type III secretion effector proteins and to examine co-factors that modulate type III secretion phenotypes.
The common theme of these projects is to better understand the mechanisms by which multidrug-resistant bacteria cause disease, resist antibiotics, and spread.
For lab information and more, see Dr. Hauser's faculty profile and lab website.
Publications
See Dr. Hauser's publications on PubMed.
Contact
Contact Dr. Hauser at 312-503-1044 or the lab at 312-503-1081.

CongCong He
Associate Professor, Cell and Developmental Biology
How autophagy (“self-eating”) carries out intracellular quality control, and regulates metabolism and behaviors in health and disease

CongCong He
Research Area(s): Cell Biology; Metabolism and Endocrinology; Neurobiology
The research in my lab is centered on intracellular quality control mediated by autophagy (“self-eating”), a lysosomal degradation pathway essential for nutrient recycling, cellular maintenance and physiological function. Autophagy is induced by stress conditions such as fasting and exercise, and allows cells to adapt to changing nutrient and energy demands through protein catabolism. Our interest focuses on the roles and mechanisms of autophagy in the regulation of metabolism and in the pathogenesis of metabolic and neurological disorders, including obesity, type 2 diabetes, neurodegeneration, and drug abuse. Malfunction of autophagy is implicated in a variety of diseases, such as metabolic disorders, neurodegeneration, cancer, infection and aging; conversely, we have shown that upregulation of autophagy mediates exercise-induced metabolic benefits and protects Alzheimer’s mice from neurodegeneration. We are also interested in demonstrating how the autophagy machinery recognizes various cargos for catabolic metabolism, including aggregate-prone proteins, secretory proteins and membrane receptors, in metabolic organs and in different neuronal cell types in the brain, and studying how such degradation leads to metabolic and behavioral alterations.
For lab information and more, see Dr. He's faculty profile and lab website.
Publications
See Dr. He's publications on PubMed.
Contact
Contact Dr. He at 312-503-3094.

Xiao-Lin He
Associate Professor, Biochemistry and Molecular Genetics
Mechanisms of signal transmission across the membrane via the cell-surface receptors

Xiao-Lin He
Research Area(s): Cancer Biology; Pharmacology
This laboratory is interested in cancer, neural development and reproduction-related structural mechanisms of how extracellular signals (e.g., growth factors, adhesion molecules and morphogens) are translated into intracellular signals by plasma membrane receptors. We use biophysical methods (crystallography, calorimetry, surface plasmon resonance, analytical ultracentrifugation, etc.) in combination with functional studies to define the physiological states and binding processes of these receptors and their complexes with ligands. Our research targets include receptor tyrosine kinases, Semaphorin and its receptors and leucine-rich-repeat-containing G-protein coupled-receptors.
For more information, visit the faculty profile of Xiaolin He, PhD.
Publications
See Dr. He's publications in PubMed.
Contact Us
Contact Dr. He at 312-503-8030 or the He Lab at 312-503-8029.

Amy B Heimberger
Professor, Neurological Surgery
New immunotherapies for patients with central nervous system (CNS) cancers informed by understanding the underlying unique immunobiology of the CNS

Amy B Heimberger
Research Area(s): Cancer Biology; Immunology; Pharmacology
Our laboratory studies the unique immunobiology of CNS tumors that informs our development of immuno-oncology therapeutics. The laboratory has been intricately involved in a wide variety of bench-to-bedside immune therapeutics, including those that developed in the laboratory and arising from our own patents. We work collaboratively with industry on their pipeline agents to clarify indications and companion biomarkers. The laboratory carries unique expertise in the investigational new drug process and window-of-opportunity clinical trials. The laboratory conducts extensive immune profiling of patient tumors including ex vivo functional assays. Our studies are directed to how various cells interact within the tumor microenvironment and the functional implications using multiplex imaging, methylation profiling, single cell sequencing and transcriptomic analysis. Areas in which we have contributed to science include the following:
EGFRvIII peptide vaccines Our laboratory co-developed with Duke University from bench-to-bedside a peptide (PEP-3-KLH/CDX-110) vaccine strategy that targets the epidermal growth factor receptor (EGFRvIII), that demonstrated induction of anti-tumor immune responses.
STAT3 mediated immune suppression and therapeutic targeting We clarified that the signal transducer and activator of transcription 3 STAT3 pathway is a key molecular hub of gliomagenesis and tumor-mediated immune suppression and conducted the preclinical development of a novel small molecule inhibitor of STAT3, WP1066, for which I hold the IND. STAT3 has been considered an “un-druggable” target and this is a first-in-man agent with specificity to STAT3. This drug has been licensed to Moleculin and is now in clinical trials.
Glioblastoma mediated mechanisms of immune suppression We have demonstrated that glioblastoma subverts the immune system to become tumor protective, especially by 1) driving tumor-associated microglia/macrophages to assist in potentiating gliomagenesis; 2) by recruitment of Tregs; 3) and by the intrinsic properties of cancer stem cells which are immunosuppressive on both adaptive and innate immunity. This investigative direction has provided potential therapeutic targets/strategies and biomarker elucidation.
miRNA and nanoparticle therapeutics The laboratory has elucidated the role of epigenetic microRNA regulation on tumor-mediated immune suppression, with an emphasis on potential translational therapeutic approaches. One of these strategies, miR-124 delivered with lipid nanoparticles to the immune compartment entered clinical trials in spontaneously arising gliomas in canines.
Immune checkpoint therapeutics and response biomarkers Given the recent FDA approvals of immune checkpoint inhibitors for malignancies, there is great enthusiasm for their use in glioblastoma. Recent work in our lab has been focused on clarification of potential response biomarkers and identification of GBM patient subsets that may benefit.
For more information, please visit Dr. Heimberger's faculty profile or the Amy Heimberger Lab page.
Publications
See Dr. Heimberger's publications on PubMed
Contact

Thomas J Hope
Professor, Cell and Developmental Biology, Obstetrics and Gynecology
Posttranscriptional regulation of intronless viral messages

Thomas J Hope
Research Area(s): Biochemistry and Structural Biology; Genetics, Epigenetics and Genomics; Microbiology; Reproductive Biology
We study the posttranscriptional regulation of intronless viral messages. Intronless messages must be efficiently processed in the absence of splicing. Therefore, intronless messages must uncouple RNA processing and export from the splicing process making a simpler model system. We are currently focused on the posttranscriptional regulatory element (PRE) of the Hepadnaviruses, including hepatitis B virus (HBV) and woodchuck hepatitis virus (WPRE). Our goal is to understand the novel mechanism of the stimulation of heterologous gene expression by the WPRE. Understanding WPRE function will allow the development of even more efficient gene expression for a variety of applications from gene therapy to large scale protein production.
Although much is known about the molecular biology of HIV, little is known about the details of interactions between the virus and cellular components such as the cytoskeleton. To gain insights into these processes we are combining the disciplines of virology and cell biology to develop the field of cellular virology. We are especially excited by new methods we have developed – such as time-lapse analysis and fluorescent tagging – that allow for HIV to be visualized in living cells.
For lab information and more, see Dr. Hope's faculty profile and lab website.
Publications
See Dr. Hope's publications on PubMed.
Contact
Contact Dr. Hope at 312-503-1360.

Lifang Hou
Professor, Preventive Medicine, Pediatrics
Environmental, genetic and epigenetic risk factors for disease

Lifang Hou
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics
Dr. Hou’s research interest lies in integrating traditional epidemiologic methods with the ever-advancing molecular techniques in multi-disciplinary research focusing on identifying key molecular markers and understanding their potential impact on disease etiology, detection and prevention.
Dr. Hou’s major research efforts to date have focused on two areas: 1) identification of risk factors that may cause chronic diseases; and 2) identification of biomarkers that serve as indicators of an individual’s past exposure to disease risk factors and/or predict future disease risks and/or prognosis. The environmental/lifestyle risk factors that Dr. Hou has studied include air pollution, pesticides, overweight, physical inactivity and reproductive factors in relation to chronic diseases. The biomarkers that Dr. Hou has investigated include genetic factors (i.e., polymorphisms, telomere length shortening, mitochondria DNA copy number variations) and epigenetic factors (i.e., DNA methylation, histone modifications and microRNA profiling). Her over-arching research goal is to understand the biological mechanisms linking environmental risk factors with subclinical or clinical disease development to ultimately lead to development of effective strategies for prevention of chronic diseases.
In addition to being a PI of several NIH funded grants, Dr. Hou is the co-director and Co-PI of the Northwestern Consortium for Early Phase Cancer Prevention Trials of the Division of Cancer Prevention (DCP) Consortia, National Cancer Institute.
For more information visit the faculty profile of Lifang Hou, MD, PhD.
Publications
See Dr. Hou's publications in PubMed.
Contact

Sui Huang
Associate Professor, Cell and Developmental Biology
Nature and function of the perinucleolar compartment (PNC), and its relationship with the malignant phenotype

Sui Huang
Research Area(s): Cancer Biology; Cell Biology
Our studies seek to understand the nature and function of a unique nuclear structure, the perinucleolar compartment (PNC), and its relationship with the malignant phenotype. Our work looks to address the function of the PNC and its significance during malignant transformation in multiple ways.
- Determine the correlation between PNCs and cancer in tissues. In collaboration with Dr. Ann Thor at Evanston Hospital, Northwestern University, we have initiated detection of PNC in breast cancer tissues.
- Investigate the function of the PNC, its relationship with the nucleolus and rRNA synthesis. We are in the process of isolating and identifying additional proteins and RNAs in the PNC.
Identification of activities taking place in the PNC will shed light on the understanding of the function of this structure and its relationship to the nucleolus and the transformed phenotype.
For lab information and more, see Dr. Huang's faculty profile.
Publications
See Dr. Huang's publications on PubMed.
Contact
Contact Dr. Huang at 312-503-4269 or the lab at 312-503-4276.

Judd F Hultquist
Associate Professor, Medicine (Infectious Diseases), Microbiology-Immunology
HIV molecular virology and translational research on pathogenic human RNA viruses

Judd F Hultquist
Research Area(s): Genetics, Epigenetics and Genomics; Immunology; Microbiology
Pathogens are molecular machines selected over millions of years of co-evolution to exploit the cellular networks of their hosts for optimal replication and survival. This reliance on and manipulation of the host manifests itself in thousands of molecular changes that influence cellular function and may result in disease. A systematic, quantitative understanding of these changes is essential for the understanding of these disease states and for the development of next-generation therapeutics. Only recently, however, have technological advances in molecular measurement and manipulation allowed for the precise, high-throughput determination of mammalian cell architecture.
Leveraging diverse expertise in primary cell models, proteomic profiling, and functional genomics, we work at the intersection of systems biology and infectious disease, defining host-pathogen interactions and their consequences for replication and pathogenesis. Through these efforts, our lab ultimately hopes to strengthen the bridge from big data to targeted discovery to clinical application for the development of personalized, host-driven therapies and the advancement of human health.
Research areas include:
- HIV-Human Network Biology
- Host Cell Regulators of HIV Latency
- RNA Virus-Host Interactions
- SARS-CoV-2 Translational Research
For lab information and more, see Dr. Hultquist's faculty profile.
Visit the Hultquist Lab Web Page or the Center for Pathogen Genomics and Microbial Evolution.
Publications
See Dr. Hultquist's publications on PubMed.
Contact
Email Dr. Hultquist
312-503-3177

Luisa Iruela-Arispe
Professor, Cell and Developmental Biology
Molecular regulation of angiogenesis and vascular homeostasis

Luisa Iruela-Arispe
Research Area(s): Cancer Biology; Cardiovascular and Renal Biology; Cell Biology
Currently, the laboratory is investigating the mechanisms behind the formation of vascular tumors and vascular anomalies. In particular, the group is interested in the identification of critical regulatory nodes that maintain vascular homeostasis and control endothelial proliferation in the context of flow. An additional focus of the lab is to dissect the molecular interactions between endothelial and tumor cells during the process of metastasis with particular emphasis on endothelial barrier.
For lab information and more, see Dr. Iruela-Arispe's faculty profile.
Publications
See Dr. Iruela-Arispe's publications on PubMed.
Contact
Email Dr. Iruela-Arispe

Fumiaki Ito
Assistant Professor, Microbiology-Immunology
Molecular basis of virus-host interactions

Fumiaki Ito
Research Area(s): Biochemistry and Structural Biology; Immunology; Microbiology
When viruses infect host cells, they establish a complex network of interactions with host factors to transform the cells into a fully habitable environment. Such interaction includes the initial attachment of the virus to the target cell surface through specific cell-entry receptors, hijacking of host cellular machinery for productive viral replication, and the evasion of host immunity through viral protein antagonists. For example, just a handful of genes encoded by human immunodeficiency virus (HIV) multitask to facilitate the infection, which involves numerous interactions with the host proteins and nucleic acids. Targeting virus-host interactions within infected cells represents an important avenue for the future development of versatile and adaptable therapeutics for many types of viral infections. To this end, we seek to gain comprehensive structural and functional information on the essential interactions between viral components and host factors.
The long-term goal in Ito laboratory is to yield atomic-level molecular insights into viral infection, host immunity, and virus-host interaction involved in various DNA and RNA virus infection by using latest cryo-electron microscopy (cryo-EM) combined with biochemical and virological functional approaches. Additionally, we aim to identify previously unrecognized host factors associated with viral proteins of interest by using a cryo-EM-based structural proteomics. We leverage the state-of-the-art cryo-EM facility at Northwestern University Feinberg School of Medicine to perform high-resolution imaging and structural analysis of virus-host complexes.
For more information, visit the faculty profile of Dr. Fumiaki Ito or the lab website
Publications
See Dr. Ito's publications in Google Scholar
Contact

Peng Ji
Professor, Pathology (Experimental Pathology), Pathology (Hematopathology)
Hematopoiesis and myeloid neoplasms

Peng Ji
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology
Our lab is interested in how cytoskeletal signaling, motor proteins and adhesion systems are integrated with chemical signaling pathways to regulate cell behavior and tissue differentiation and disease. The Ji lab studies small G proteins and downstream actin regulatory effectors that participate in enucleation during red cell development.
At the level of the nucleus, the Ji laboratory studies genes involved in erythroid lineage commitment, chromatin condensation and enucleation towards understanding how congenital red cell disorders and leukemia develop.
For more information, visit the faculty profile of Peng Ji, MD, PhD or the lab website
Publications
See Dr. Ji's publications in PubMed.
Contact

Bin Jiang
Research Assistant Professor, Surgery (Vascular Surgery)
Research Assistant Professor, McCormick School of Engineering
Vascular disease modeling, tissue engineering and regenerative medicine

Bin Jiang
Research Area(s): Cardiovascular and Renal Biology; Developmental and Stem Cell Biology; Metabolism and Endocrinology
The Jiang laboratory is an interdisciplinary research program of vascular surgery and biomedical engineering. The primary focus of our work is on vascular repair and regeneration for a variety of vascular diseases and conditions. We use a combination of innovative technologies, including induced pluripotent stem cells (iPSCs), biomaterials, and non-invasive imaging to develop patient-specific, tissue engineered vascular constructs. Currently, the laboratory is investigating the mechanisms behind vascular calcification and abdominal aortic aneurysm with novel disease models in vitro and in vivo. Additionally, we explore the role of microenvironment in the differentiation of vascular cell phenotypes in health and disease. Ultimately, the scientific discoveries and engineering solutions developed by our research program will benefit patients suffering from vascular diseases.
For more information, visit Dr. Jiang's faculty profile page or the Jiang lab website
Publications
See Dr. Jiang's publications.
Contact

Sumanas W Jordan
Assistant Professor, Surgery (Plastic Surgery)
Designing electroconductive scaffolds to enhance tissue reinnervation

Sumanas W Jordan
Research Area(s): Developmental and Stem Cell Biology; Neurobiology
Our lab takes inspiration from difficult problems encountered in plastic surgery, specifically peripheral nerve regeneration and denervation injury, lymphedema, and vascularized composite allotransplantation. The overarching theme of the lab is electroconductive biomaterial design for regeneration and disease state modeling. Neurons and myocytes are classic examples of electroactive cells that respond to electrical fields and stimulation. However, other cell types such as endothelium and smooth muscle likely also exhibit responsiveness to electrical fields and stimulation. Early projects are focused on the development of graphene-PLGA based scaffolds for nerve and muscle co-regeneration and the role of decellularized tissue matrices in material design.
For lab information and more, visit Dr. Jordan's faculty profile
Publications
See Dr. Jordan's publications on PubMed.
Contact
Contact Dr. Jordan

Robert G Kalb
Professor, Neurology (Neuromuscular Disease)
Activity-dependent development of circuits in the central nervous system and neurodegenerative diseases

Robert G Kalb
Research Area(s): Neurobiology
Glutamatergic synapses that include the GluA1 subunit have a privileged role in activity-dependent brain development and this is driven, in part, through the assembly of a large multi-protein complex in the post-synaptic density. A critical molecular component of this complex is the scaffolding protein SAP97. Among the >90 known SAP97 binding partners we have determined that a novel protein called CRIPT plays an essential role in this process. Humans with mutations in CRIPT have a severe developmental brain disorder. We are using a variety of approaches to understand the mechanisms by which CRIPT controls synapse biology and dendrite growth including: 1) study of a conditional knock-out (cKO) mouse that we built, 2) super-resolution imaging of glutamate receptors/TARPs/MAGUKs using neurons derived from patient iPS cells and 3) electrophysiology of dissociated neurons and hippocampal slices from the cKO mice. Insight into the molecular logic of SAP97/CRIPT function will have implications for childhood maladies such as intellectual disability and autism/autism-spectrum disorders.
In our studies of adult onset neurodegenerative diseases such as Amyotrophic Lateral Sclerosis and Frontotemporal Dementia we find evidence for maladaptive changes in cellular intermediary metabolism and proteostasis. Ongoing metabolomics interrogations are uncovering why changes in fuel utilization are toxic and discovering new targets for therapeutic intervention. The relationship between altered metabolism and perturbed protein homeostasis is also an area of intense interest with special focus on a proteasome adaptor protein called RAD23. Our experimental platforms are: 1) cells from patients with various genetic abnormalities (i.e., mutations in C9orf72, TDP43, etc.) differentiated into neurons, 2) C.elegans, and 3) primary rat/mouse neurons. Targeting proximal events in neurodegenerative diseases will lead to novel therapeutic approaches.
For lab information and more, visit Dr. Kalb's faculty profile or the Kalb Lab website.
Publications
See Dr. Kalb's publications on PubMed.
Contact
Contact Dr. Kalb at 312-503-5358

Atsushi Kato
Associate Professor, Medicine (Allergy and Immunology), Otolaryngology - Head and Neck Surgery
Mechanisms of initiation and amplification of type 2 inflammation in airway inflammatory diseases in humans

Atsushi Kato
Research Area(s): Immunology
The Kato Lab primarily focuses on the mechanism of type 2 inflammation in airway inflammatory diseases in humans. Currently, we use chronic rhinosinusitis (CRS) with nasal polyps (CRSwNP) as a model disease of type 2 inflammation. CRS is an inflammation of the nose and sinuses that blocks the air passages, causes headache and leads to loss of sleep, depression and reduced quality of life. CRSwNP which is a severe case of CRS, is well characterized by tissue eosinophilia with high levels of type 2 cytokines including IL-5 and IL-13. However, the mechanisms of type 2 inflammation in nasal polyps are still not well understood. The Kato Lab is currently focused on an epithelial-derived cytokine, TSLP (thymic stromal lymphopoietin), that is an IL-7-like cytokine molecule and is now recognized as an important regulator of type 2 inflammation in nasal polyps. We recently identified that TSLP is highly up-regulated in nasal polyps. TSLP is known to directly and indirectly induce type 2 inflammation via the activation of dendritic cells, Th2 cells, group 2 innate lymphoid cells (ILC2s) and mast cells which are all elevated in nasal polyps. My laboratory is currently investigating the role of Th2 cells, ILC2s, mast cells and dendritic cells in the amplification of type 2 inflammation and how TSLP contributes to type 2 inflammation through these cell types in nasal polyps. In contrast to CRSwNP, inflammatory patterns in non-polypoid CRS (CRSsNP) are much less understood. Recently, the PI’s laboratory fully characterized patterns of inflammatory cytokines in the nasal mucosa of control subjects and patients with CRS. We found that CRSsNP displays heterogenous inflammation and this heterogeneity in CRSsNP might be responsible for the inconsistency of results in CRSsNP-related studies. We are also currently working on understanding the effect of inflammatory endotypes on clinical phenotypes in CRSsNP.
For more information, view the faculty profile for Atsushi Kato, PhD.
Publications
See Dr. Kato's publications in PubMed.
Contact

Jennifer A Kearney
Associate Professor, Pharmacology
Genetic basis of epilepsy

Jennifer A Kearney
Research Area(s): Genetics, Epigenetics and Genomics; Neurobiology; Pharmacology

Research Description
My research program is focused on studying the genetic basis of epilepsy, a common neurological disorder that affects approximately 1% of the population. Epilepsy is thought to have a genetic basis in approximately two-thirds of patients, including a small fraction caused by single gene mutations. Many genes responsible for rare, monogenic epilepsy have been identified. The genes identified are components of neuronal signaling, including voltage-gated ion channels, neurotransmitter receptors, ion-channel associated proteins and synaptic proteins. We use mouse models with mutations in ion channel genes to understand the underlying molecular basis of epilepsy and to identify modifier genes that influence phenotype severity by modifying disease risk. Identifying genes that influence epilepsy risk improves our understanding of the underlying pathophysiology and suggests novel targets for therapeutic intervention.
For lab information and more, see Dr. Kearney's faculty profile.
Publications
See Dr. Kearney's publications on PubMed.
Contact
Contact Dr. Kearney at 312-503-4894.

Neil Lindstrom Kelleher
Professor, Medicine (Hematology and Oncology), Biochemistry and Molecular Genetics
Professor, Weinberg College of Arts and Sciences
Top down proteomics, natural products discovery and biosynthesis and chromatin oncobiology and DNA-damage.

Neil Lindstrom Kelleher
Research Area(s): Biochemistry and Structural Biology; Genetics, Epigenetics and Genomics
The main focus for our Top Down Proteomics subgroup is to push the limits for whole proteome analysis of mammalian cells, striving for a future in which Top Down analysis rivals that of Bottom Up in the number of protein identifications per run. Recently, we have seen progress toward this very goal with the introduction of a separation platform specifically designed to minimize the most common problem in Top Down Proteomics, intact protein separations. This platform effectively reduces sample complexity and separates proteins depending on size, resulting in an opportunity for the scientist to select the optimal analysis method for the sample.
Our Natural Products subgroup is focused on the discovery and biosynthesis of novel natural products. Developments from this subgroup include the introduction of the PrISM platform, geared towards the identification of natural products synthesized by nonribosomal peptide synthetases (NRPSs) and polyketide synthases (PKSs) without prior knowledge of a gene sequence. This is made possible by our ability to detect a phosphopantetheinyl (Ppant) ejection marker ion for NRPS/PKS thiolation domains. We also work in collaboration with groups from other universities to provide mass spectrometry analysis of novel biochemical systems.
We also have a long-standing interest in histone analysis. Our Chromatin Oncobiology and DNA-Damage subgroup continues to dig deeper into the "histone code", a complex mixture of post-translational modifications that together determine a host of cellular processes. We are interested in visualizing dynamic histone PTM changes simultaneously on multiple sites. Through application of technology developed in our Top Down Proteomics subgroup, we are able to apply "Precision Proteomics" to histone analysis.
For more information, visit the Kelleher Lab Web Page or see Dr. Kelleher's faculty profile.
Publications
View lab publications via PubMed.
Contact Us
Contact Dr. Kelleher or the Kelleher Lab at 847-467-1086 or 847-467-4362.

Shana O Kelley
Professor, Biochemistry and Molecular Genetics
Professor, Weinberg College of Arts and Sciences
New Technologies for Disease Biology

Shana O Kelley
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Pharmacology
The Kelley lab utilizes an interdisciplinary approach that integrates nanoscience, bioanalytical science and engineering, focusing on high-throughput single-cell profiling and the application of new technology platforms of the characterization of pathways relevant to cancer progression and treatment.
For more information, see Dr. Kelley's faculty profile or the Kelley lab website.
Publications
See Dr. Kelley's publications on PubMed
Contact

John A Kessler
Professor, Neurology (Comprehensive Neurology), Pharmacology
Biology of neural stem cells and growth factors and their potential for regenerating the damaged or diseased nervous system

John A Kessler
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Developmental and Stem Cell Biology; Neurobiology
The Kessler laboratory focuses on the biology of neural stem cells and growth factors and their potential for regenerating the damaged or diseased nervous system. A major interest of the laboratory has been the role of bone morphogenetic protein (BMP) signaling in both neurogenesis and gliogenesis and in regulating cell numbers in the developing nervous system. Both multipotent neural stem cells and pluripotent embryonic stem cells are studied in the laboratory. Recent efforts have emphasized studies of human embryonic stem cells (hESC) and human induced pluripotent stem cells (hIPSC). The Kessler lab oversees the Northwestern University ESC and IPSC core and multiple collaborators use the facility. In addition to the studies of the basic biology of stem cells, the laboratory seeks to develop techniques for promoting neural repair in animal models of spinal cord injury and stroke. In particular, the lab is examining how stem cells and self-assembling peptide amphiphiles can be used together to accomplish neural repair. The lab is also using hIPSCs to model Alzheimer’s disease and other disorders.
For more information see the faculty profile of John A. Kessler, MD.
Publications
View Dr. Kessler's full list of publications in PubMed.
Contact

Dong-Hyun Kim
Professor, Radiology (Basic and Translational Radiology Research)
Professor, McCormick School of Engineering
Therapeutic micro/nanoparticles and their hybrid derivatives for treatment of various types of cancer

Dong-Hyun Kim
Research Area(s): Biochemistry and Structural Biology; Cancer Biology
Image-guided medicine is rapidly growing to improve treatment regimens as advancing medical imaging, including magnetic resonance imaging (MRI), computed tomography (CT), radiography, ultrasound, positron emission tomography (PET), and single photon emission computed tomography (SPECT). A combination of modern nanoplatforms with high performance in imaging and therapeutics may be critical to improve medical outcomes. One of emerging fields is the image guided therapy using various nanoparticles. Those are including basic bench, preclinical in vitro/in vivo and clinical researches combining synthesis of multifunctional nanoparticle and tracking/navigation tools to improve accuracy and outcomes of the therapeutics. Most of the emerging interventional technique such as heat activated targeted drug delivery, image guided ablation (microwave or HIFU), percutaneous injection gene/bacteria therapy, transcatheter treatments for tumor specific local therapy, serial biopsy, thrombolytic therapy, and so on, can be combined with nanotechnology in clinic. Careful design/selection/synthesis of multifunctional imaging/therapeutic nanomaterials with therapeutic agents will be critical for the translational optimization these new image guided medicine techniques. The DHKIM Lab for Biomaterials of Image Guided NanoMedicine has focused on developing various therapeutic/imaging carriers for the treatment of various cancers. Micro/Nanoparticles and their hybrid derivatives have been exploited as vectors for drug/therapeutic delivery and molecular imaging agents of MRI, CT, ultrasound and luminescent/fluorescents. We are working closely with clinicians, medical scientists, biologist and imaging professionals to translate new therapeutic approaches using multifunctional carriers and diagnostic imaging technique to the clinical setting.
For more information, please see Dr. Kim's faculty profile or visit the Kim Lab Website
Publications
See Dr. Kim's publications in PubMed.
Contact
Email Dr. Kim
Phone Kim lab 312-926-3279

Ji-Yong Julie Kim
Professor, Obstetrics and Gynecology (Reproductive Science in Medicine)
Molecular mechanisms of uterine disease development and growth

Ji-Yong Julie Kim
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology; Reproductive Biology
Our research focuses on the molecular mechanisms that underlie the development and growth of endometrial cancer, endometriosis, and fibroids. We aim to identify novel therapeutic targets for these diseases by investigating the role of sex hormones, genetics, race, and risk factors such as obesity. To study disease initiation and progression, we require appropriate model systems that accurately reflect the in vivo situation. Therefore, we are invested in building physiological models of the endometrium and myometrium. We recognize the importance of tissue architecture and have generated organoids and spheroids of the endometrium and myometrium. We also study the contribution of genetics by using iPSCs to study endometriosis and to recreate the disease in vitro. Our models will provide a system to understand human physiology and can serve as a screening tool for drug discovery.
For more information, please see Dr. Kim's faculty profile or the Kim Lab website.
Publications
See Dr. Kim's publications in PubMed.
Contact
Contact Dr. Kim at 312-503-5377 or the Kim Lab at 312-503-4762.

Evangelos Kiskinis
Associate Professor, Neurology (Neuromuscular Disease), Neuroscience
Molecular mechanisms that give rise to neurological diseases using human stem cell-derived neuronal subtypes

Evangelos Kiskinis
Research Area(s): Neurobiology
The broad objective of our laboratory is to understand the nature of the degenerative processes that drive neurological disease in human patients. We are primarily interested in Amyotrophic Lateral Sclerosis (ALS), Epileptic Syndromes as well as the age-associated changes that take place in the Central Nervous System (CNS). We pursue this objective by creating in vitro models of disease. We utilize patient-specific induced pluripotent stem cells and direct reprogramming methods to generate different neuronal subtypes of the human CNS. We then study these cells by a combination of genomic approaches and functional physiological assays. Our hope is that these disease model systems will help us identify points of effective and targeted therapeutic intervention.
For more information view the faculty profile of Evangelos Kiskinis, PhD, or the Evangelos Kiskinis Lab site.
Publications
View Dr. Kiskinis' publications at PubMed.
Contact Information
Hiroaki Kiyokawa
Professor, Pharmacology, Pathology
Roles of cell cycle-regulatory proteins in differentiation, senescence and tumorigenesis and cell cycle control in endocrine and reproductive organs
Hiroaki Kiyokawa
Research Area(s): Cancer Biology; Reproductive Biology
We are interested in the basic mechanisms of cell cycle control, cellular senescence/immortalization and malignant transformation, with a focus on protein regulation by ubiquitination. We previously demonstrated that cell cycle regulators such as p27Kip1, CDK4 and CDC25A play highly tissue-specific roles in development and oncogenesis. Ubiquitination, the covalent modification of substrate proteins with the small 76-residue protein ubiquitin, exerts diverse regulation of the fate of substrates, including the cell cycle regulators, e.g, promoting proteolysis, altering subcellular localization and modulating enzymatic activities. Our current research is aimed at revealing novel functions of ubiquitination enzymes and their substrates in development and cancer, which is expected to identify new therapeutic targets against human diseases. The laboratory uses a combination of protein engineering, proteomics, bioinformatics, cell biological techniques such as time-lapse microscopy and 3-D culture and genetically engineered mouse models. Keywords: cell cycle, ubiquitin, ubiquitination, cancer initiation, cancer progression, knockout mice, transgenic mice, breast cancer, cyclin, diabetes, pituitary, development.
Recent Findings
- There is a unique regulation of cell cycle progression in neuroendocrine tissues such as pancreatic islets and pituitary glands of CDK4-null mice; we have shown that in this particular type of cell cycle, Cdk4 plays an indispensable and rate-limiting role
- CDC25A phosphatase, which activates CDK2 and CDK1, is an oncogene that plays a rate-limiting role in initiation and progression of various tumors, including breast cancer
Current Projects
We are currently investigating roles of the cell cycle machinery in differentiation, tumorigenesis and apoptosis, by combinations of mouse models and molecular analyses.
For lab information and more, see Dr. Kiyokawa’s faculty profile.
Publications
See Dr. Kiyokawa's publications on PubMed.
Contact
Contact Dr. Kiyokawa at 312-503-0699.

David J Klumpp
Professor, Urology, Microbiology-Immunology
Molecular Mechanisms Of Bladder Inflammation and Pelvic Pain

David J Klumpp
Research Area(s): Immunology; Microbiology; Pharmacology
Our laboratory employs state-of-the-art cell culture and animal models to pursue multi-disciplinary projects in bacterial pathogenesis and neuro-immune interactions in a crippling pain syndrome. A key to our success is the rich training environment resulting from the close collaboration between clinical and basic scientists.
Urinary tract infection (UTI) is both a major medical issue and a fascinating problem of bacterial pathogenesis. We investigate all aspects of host pathogen interactions, from the immediate biochemical signaling evoked in bladder cells, to the associated inflammation, to the development of adaptive immune responses. We recently identified a novel signaling response of bladder cells induced by binding of uropathogenic E. coli (UPEC) that mediates the mutually exclusive processes of epithelial cell apoptosis and bacterial invasion of bladder epithelial cells; how this occurs is an active area of study. We have also identified a candidate live-attenuated UTI vaccine based on a UPEC mutant. We find that the UPEC mutant vaccine induces protective responses 100-fold greater than wild type UPEC. We are now determining the mechanism of this enhanced response by testing the hypothesis that the UPEC mutant skews the normal immune response and thus generates a more effective immunity.
Although pelvic pain can result from acute infection, interstitial cystitis (IC) is a debilitating chronic pelvic pain syndrome of unknown origin that is often considered a chronic bladder inflammation. We utilize a herpesvirus to induce an IC-like condition in mice Using this model, we have identified the mechanisms that result in both bladder pathophysiology and pelvic pain. Interestingly, while both pain and bladder damage require mast cell activation, pelvic pain results from the release of mast cell histamine, whereas bladder pathology is driven by mast cell release of tumor necrosis alpha (TNF). Current studies include the genetic basis of pain susceptibility, the spinal regulation of histamine and TNF release and the viral basis of pain. In addition, we recently were awarded a prestigious NIH center grant to study pelvic pain syndromes. The center award will extend our bladder pelvic pain studies to prostate- and bowel-associated pelvic pain and determine the mechanisms of pelvic organ crosstalk and signal integration in the spinal cord in mice. Our center collaborators will examine cortical and cognitive changes in pelvic pain patients using a combination of functional MRI and behavioral tests and develop novel quality-of-life tests to characterize pelvic pain non-invasively within populations. Thus, this center will illuminate pelvic pain mechanisms in clinical, epidemiologic and basic animal studies.
For more information, visit Dr. Klumpp's faculty profile.
Publications
See Dr. Klumpp's publications in PubMed.
Contact

Igor J Koralnik
Professor, Neurology (Neuro-infectious Disease and Global Neurology)
Viral effects on the nervous system

Igor J Koralnik
Research Area(s): Microbiology; Neurobiology
The laboratory of Igor Koralnik, MD, studies how viruses affect the nervous system. These include SARS-CoV-2 in patients with COVID-19, HIV in patients with substance use, as well as the entire virome in patients with neurodegenerative diseases. In addition, the Koralnik Lab is involved in global neurology research with international partners.
For more information and publications see the faculty profile of Igor Koralnik, visit the Koralnik Lab or the Program for Global Neurology.
Publications
See Dr. Koralnik's publications on PubMed.
Contact

Dimitri Krainc
Professor, Neurology (Movement Disorders), Neuroscience, Neurological Surgery
Professor, Weinberg College of Arts and Sciences
Mechanisms of neuronal dysfunction in neurodegenerative disorders that affect children and adults

Dimitri Krainc
Research Area(s): Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Pharmacology
The overarching goal of my laboratory is to study rare diseases such Huntington’s and genetic forms of Parkinson’s disease, as a window to understanding neurodegeneration across the lifespan. More recently, we have focused on rare lysosomal diseases such as Gaucher’s in order to identify specific targets and mechanisms that contribute to neurodegeneration in Parkinson’s and related synucleinopathies. It is expected that such defined targets will facilitate mechanism-based development of targeted therapies for children with neuronopathis Gaucher’s disease as well as adult-onset synucleinopathies such as Parkinson’s disease. To validate and study these targets and novel therapies in human neurons, we have utilized induced pluripotent stem cells (iPS) generated by reprogramming of patient-specific skin fibroblasts. These iPS cells are differentiated into specific neuronal subtypes in order to characterize the contribution of genetic, epigenetic and environmental factors to disease mechanisms and to test novel therapeutic approaches.
For more information see the faculty profile of Dimitri Krainc or visit the Krainc Lab website.
Recent Publications
View Dr. Krainc's full list of publications in PubMed.
Contact information
Dimitri Krainc, MD, PhD
Ward Professor and Chairman
312-503-3936

Fei Li Kuang
Assistant Professor, Medicine (Allergy and Immunology)
Impact of Eosinophils in Allergic Diseases

Fei Li Kuang
Research Area(s): Immunology
The Kuang Lab is a translational research group that investigates the roles of eosinophils and lymphocytes, two different white blood cells, in a variety of allergic and eosinophilic disorders such as asthma, medication-induced eosinophilia vs. DRESS, EGID and hypereosinophilic syndromes (HES). We create well-characterized human patient cohorts and collect both clinical data and biological samples from human research subjects, with the goal of discovering unique blood eosinophil and T cell signatures in EGID as compared to other disorders. We then utilize cell culture systems to dissect out the molecular pathways implicated in disease pathogenesis. Our findings could transform how different types of allergic and eosinophilic diseases (e.g. allergic asthma) are understood and managed. These studies are particularly timely as more patients are placed on drugs that remove eosinophils or target T cells.
Publications
View Dr. Kuang's publications on PubMed.
For more information, visit the faculty profile for Fei Li Kuang, MD, PhD.
Contact Us
Contact Dr. Kuang at 312-503-0077

Tsutomu Kume
Professor, Medicine (Cardiology), Pharmacology, Ophthalmology
Cardiovascular development, cardiovascular stem/progenitor cells and angiogenesis

Tsutomu Kume
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Developmental and Stem Cell Biology
Cardiovascular development is at the center of all the work that goes on in the Kume lab. The cardiovascular system is the first functional unit to form during embryonic development and is essential for the growth and nurturing of other developing organs. Failure to form the cardiovascular system often leads to embryonic lethality and inherited disorders of the cardiovascular system are quite common in humans. The causes and underlying developmental mechanisms of these disorders, however, are poorly understood. A particular emphasis in our laboratory has recently been the study of cardiovascular signaling pathways and transcriptional regulation in physiological and pathological settings using mice as animal models, as well as embryonic stem (ES) cells as an in vitro differentiation system. The ultimate goal of our research is to provide new insights into the mechanisms that lead to the development of therapeutic strategies designed to treat clinically relevant conditions of pathological neovascularization.
Publications
View Dr. Kume's publications on PubMed.
For more information, visit the faculty profile for Tsutomu Kume, PhD.
Contact Us
Contact Dr. Kume at 312-503-0623 or the Kume Lab at 312-503-3008.

Laimonis A Laimins
Professor, Microbiology-Immunology
Molecular biology of human papillomaviruses (HPV) and their association with cervical cancer

Laimonis A Laimins
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics; Microbiology; Reproductive Biology
Our efforts are divided into two main categories:
- An examination of how the viral oncoproteins E6 and E7 contribute to the development of malignancy
- Studies on the mechanisms controlling the viral life cycle in differentiating epithelia
More than 100 distinct types of human papillomavirus have been identified and approximately one-third specifically target squamous epithelial cells in the genital tract. Of these genital papillomaviruses, a subset including types 16,18 and 31 have been shown to be the etiological agents of most cervical cancers.
One of our goals is to understand why infection by specific HPV types contributes to the development of malignancy. For these studies we have examined the interaction of the oncoproteins E6 and E7 with cellular proteins. In particular, E6 binds the p53 protein and facilitates its degradation by a ubiquitin-mediated pathway. It also activates telomerase as well as associates with PDZ-domain containing proteins. The interactions of the E6 and E7 proteins with these cellular proteins are being examined at both the biochemical and genetic level.
In examining the papillomavirus life cycle, we have used organotypic tissue culture systems to faithfully reproduce the differentiation program of epithelial cells in the laboratory. Using this system, the viral life cycle has been duplicated. We are studying the mechanisms that regulate viral DNA replication, cell entry, immune evasion and gene expression. These studies should provide insight into viral pathogenesis as well as the mechanisms regulating epithelial differentiation.
For lab information and more, see Dr. Laimins' faculty profile and lab website.
Publications
See Dr. Laimins' publications on PubMed.
Contact
Contact Dr. Laimins at 312-503-0648 or the lab at 312-503-0650.

Monica M Laronda
Associate Professor, Pediatrics (Endocrinology), Obstetrics and Gynecology
Pediatric fertility and hormone preservation and restoration

Monica M Laronda
Research Area(s): Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Reproductive Biology
Our research addresses fundamental regenerative medicine questions through the lens of reproductive biology. The main objective of our lab is to develop a patient-specific ovarian follicle niche that will support systemic endocrine function and fertility in women and girls who were sterilized by cancer treatments, have disorders of sex development or were exposed to other factors that could result in premature ovarian failure or sex hormone insufficiency. This research is a part of the Ann & Robert H Lurie Children’s Hospital Fertility and Hormone Preservation and Restoration Program that bridges basic science, translational research and clinical practice.
For more information see Dr. Laronda's faculty profile, the Laronda lab website, the Laronda lab at Lurie Children's Hospital, or the Fertility and Hormone Preservation and Restoration Program.
Publications
See Dr. Laronda's publications in PubMed.
Contact

Shannon M Lauberth
Associate Professor, Biochemistry and Molecular Genetics
Decoding the cancer epigenome

Shannon M Lauberth
Research Area(s): Cancer Biology; Cell Biology; Genetics, Epigenetics and Genomics; Immunology
The Lauberth Lab is located in the department of Biochemistry and Molecular Genetics at Northwestern University Feinberg School of Medicine and is located in the Simpson Querrey Biomedical Research Center on Northwestern’s campus in downtown Chicago.
Our primary research focus is to make advances in basic and translational studies in the cancer epigenetics field. The laboratory utilizes a combination of approaches that include cutting-edge high-throughput NGS technologies, state of the art microscopy techniques, quantitative proteomics, biochemistry, and cell-based/genetic assays. We also employ powerful cell-free assays that fully reconstitute transcription on chromatin templates and are powerful in discerning direct (causal) effects of epigenetic and transcriptional regulatory mechanisms.
Current Projects
The general transcription machinery functions as signal transducers
Through an exciting collaboration with Nullin Divecha’s group (University of Southampton), we discovered a new role of the basal transcription TFIID complex component, TAF3 in transducing nuclear phosphoinositide signals into gene expression changes that are required to build muscle tissue. This work was published in Molecular Cell.
p53 mutant enhancer selection and regulation imparts transcriptional plasticity to cancer cells in response to chronic immune signaling
My lab has made important contributions in revealing mechanistic insights into how chronic immune signaling drives alterations in the cancer cell transcriptome. We have demonstrated a functional crosstalk between the second most frequently identified p53 mutant and the master proinflammatory regulator NFkB that shapes an active enhancer landscape to reprogram the cancer cell transcriptome in response to chronic TNFa signaling. These studies also revealed insights into mutant p53-dependent gene regulation by describing new ways in which mutant p53 is recruited to specific classes of enhancers through various binding partners since mutant p53 does not directly engage with DNA. This new mechanism underlying the tumor-promoting roles of mutant p53 was published in Nature Communications.
Mutant p53 Co-Opts Chromatin Pathways at Enhancers to Drive Cancer Cell Growth
Our studies have advanced our understanding of cancer epigenetics by establishing an interplay between p53 mutants and chromatin regulators that leads to aberrant enhancer and gene activation in human cancer. Specifically, through global analyses, we demonstrated a requirement for mutant p53 in regulating the prominent histone mark, monomethylation of histone H3 lysine 4 (H3K4me1) that demarcates enhancer regions. This is an important finding that provides new mechanistic insights into how H3K4me1 levels are altered, which is a frequent event in various human cancers. Also, by implementing cell-based and our unique cell-free assays, we revealed mechanistic insights into the cooperativity between epigenetic regulators, MLL4 and the histone acetyltransferase p300 in promoting joint epigenetic aberrations that support enhanced transcriptional activation by mutant p53. These important findings were published in the Journal of Biological Chemistry.
eRNAs regulate the expression of tumor promoting genes
Our recent research efforts have resulted in new insights into the functions of eRNAs, which is particularly significant since these findings are among the few studies to date that have identified roles for these noncoding transcripts. Through global profiling analyses, we identified eRNAs that are synthesized by various p53 mutants, and by depleting several of these eRNAs, we identified their direct roles in the regulation of tumor promoting gene expression. We also revealed that these eRNAs function through the chromatin recognition bromodomains (BDs) of the extra-terminal motif (BET) family member BRD4. We further characterized a mechanism in which eRNAs are required to increase BRD4 binding to acetylated histones and promote enhanced BRD4 and RNAPII recruitment at specific enhancers that, in turn augments enhancer and tumor promoting gene activation. We also extended the implications of our findings by revealing that the BDs of all BET family members and several non-BET family members also directly interact with eRNAs. These findings provide a previously unrecognized convergence between eRNAs and histone posttranslational modifications in regulating the binding of chromatin reader proteins and highlight a mechanism in which eRNAs play a direct role in gene regulation by modulating chromatin interactions and transcription functions of BRD4. A manuscript describing these findings has been published in Nature Structural & Molecular Biology (NSMB).
We have also published several reviews on enhancer regulation and function and the mechanisms underlying eRNA functions that you can find by viewing our publications.
For more information, please see Dr. Lauberth's faculty profile and laboratory website.
Publications
View all of Dr. Lauberth's publications on PubMed.
Contact
Contact Dr. Lauberth at 312-503-4780.
Lab Contact Information
Northwestern University Feinberg School of Medicine
Simpson Querrey 7-400B
303 E. Superior Street
Chicago, IL 60611

Jeremy A Lavine
Assistant Professor, Ophthalmology, Medicine (Rheumatology)
New therapeutics for ocular angiogenesis independent of vascular endothelial growth factor (VEGF)

Jeremy A Lavine
Research Area(s): Cell Biology; Immunology; Neurobiology
My research lab focuses on translational, basic science projects that aim to develop new therapeutics for ocular angiogenesis independent of vascular endothelial growth factor (VEGF). Neovascular age-related macular degeneration (nAMD) is the leading cause of visual impairment in the developed world. Currently, humanized anti-VEGF antibodies are the gold standard for the treatment of nAMD. Patients currently undergo frequent (up to monthly) injections of anti-VEGF antibodies into the vitreous cavity. The average patient achieves 1-2 lines of visual acuity gain, but 15% of patients still lose vision despite maximal anti-VEGF therapy. Although 15% appears small, given the high prevalence of nAMD, this amounts to 2.5 million patients worldwide. For these poorly responsive patients, there is a clear unmet need for alternative, VEGF-independent therapeutic options.
Macrophage recruitment is central in nAMD pathogenesis. Choroidal neovascularization (CNV) is the pathological hallmark of nAMD. In human histopathology studies of excised CNV membranes, macrophages are readily apparent. In mice, nAMD is modeled by laser-induced injury, which causes CNV membrane formation. Laser-induced CNV formation is robustly inhibited by chemical or genetic macrophage depletion. Based upon these accepted dogma, intravitreal steroids were attempted for nAMD treatment and are unfortunately ineffective. I hypothesize that steroids anti-inflammatory properties are too broad and specific anti-macrophage therapies are necessary. Furthermore, macrophages are highly plastic and heterogenous populations, including pro-inflammatory, pro-restorative, pro-fibrotic, and pro-angiogenic subtypes. My lab’s focus is to identify macrophage heterogeneity in CNV, delineate pro-angiogenic macrophage subtypes, and attempt to develop therapies against pro-angiogenic macrophages for nAMD.
For more information, visit the faculty profile for Dr. Lavine or the Lavine lab website.
Publications
See Dr. Lavine's publications on PubMed.
Contact
Lab Phone: 312-503-0487

I. Caroline Le Poole
Professor, Dermatology, Microbiology-Immunology
Immune recognition of melanoma-associated antigens, aiming to develop immunotherapeutics for benign and malignant disease

I. Caroline Le Poole
Research Area(s): Cancer Biology; Cell Biology; Immunology; Microbiology
Role for HSP70i in driving autoimmune responses. This stress protein is included as a component of some anti-tumor vaccines based on its chaperone and adjuvant functions. In human samples, we found that HSP70i is upregulated in vitiligo skin, that HSP70i can associate with melanosomes under stress and that the heat shock protein is increasingly secreted by vitiligo melanocytes.
Development of an immunotherapeutic vaccine for patients with lymphangioleiomyomatosis (LAM). This devastating disease involves development of slow growing tumors in the lungs of female patients with mutations in TSC1 or TSC2. As tumor cells of smooth muscle cell origin transdifferentiate to express melanoma associated antigens, we propose to target melanosomal antigens using T cell receptor transgenic, autologous T cells.
Development of a chemopreventive approach towards melanoma. The concept carries similarity to ‘elective surgery’ available for some other cancers. This strategy is further supported by the immune response that is indirectly elicited by dying melanocytes targeted by topically applied bleaching phenols. We have contributed to elucidating the mechanism by which phenolic agents can induce melanocyte death, expanded the arsenal of reagents to include those which specifically target the stem cell population and studied the immune response that follows.
Restoring tolerance in newly developed mouse models of autoimmune vitiligo. Contrary to the existing problem of overzealous regulatory responses that interfere with anti-tumor immunity, patients with autoimmune disease activity generally lack effective regulatory responses. So whereas anti-CD25 therapy to deplete Tregs is popular as a pretreatment for patients undergoing anti-tumor immunotherapy, the opposite holds true in autoimmune disease. We propose to manipulate Treg homing in particular, and recently demonstrated elevated CCR4 expression in circulating Treg from melanoma patients whereas its ligand is highly expressed in tumors; the opposite holds true in vitiligo.
For more information, see the faculty profile for Dr. Le Poole, PhD
Publications
See Dr. Le Poole's publications on PubMed
Contact

Catalina Lee Chang
Assistant Professor, Neurological Surgery
Translational neuro-immuno-oncology

Catalina Lee Chang
Research Area(s): Cancer Biology; Immunology; Microbiology; Neurobiology
The Lee-Chang is a translational neuro-immuno-oncology lab focusing on understanding the interaction between CNS tumors and the immune system. We study how brain tumors suppress the immune response to leverage current and novel immunotherapies. We are particularly interested in developing genetically engineered B-cell-based therapies that promote an anti-tumoral cytotoxic immune response.
For more information, please see Dr. Lee-Chang's faculty profile.
Publications
See Dr. Lee-Chang's publications on PubMed.
Contact

Liming Li
Associate Professor, Biochemistry and Molecular Genetics
Structural properties of prion proteins using yeast as a model organism

Liming Li
Research Area(s): Biochemistry and Structural Biology; Neurobiology
Prion diseases belong to a class of fatal, infectious neurodegenerative diseases known as transmissible spongiform encephalopathies (TSEs), including the bovine spongiform encephalopathies (BSE or mad cow disease) in cattle and Creutzfeldt-Jakob disease (CJD) in human. It is generally accepted that the infectious agent of prion disease is a normal host protein (PrPC) that has adopted a pathogenic conformation that is infectious (PrPSc). Remarkably, there are several atypical yeast proteins capable of existing in multiple stable conformations, each of which is associated with distinct phenotypes. Intriguingly, some of the conformations are able to self-propagate and are “infectious.” They are thus referred to as yeast prions. Our laboratory is interested in study this fascinating prion phenomenon using yeast as a model organism. Yeast offers a powerful system that is amenable to biochemical, cell biological and genetic manipulations. We want to obtain information on the structural properties of yeast prions, their mutual interactions and their interactions with other cellular factors, particularly, with molecular chaperones. We have recently discovered that the yeast heat-shock transcription factor (HSF), a master regulator of molecular chaperones’ production, plays an important role in governing the de novo formation and “strain” determination of yeast prion [PSI+]. We are working toward to identify novel cellular factors that are HSF targets and important for yeast prion formation and inheritance. The function of HSF is evolutionally conserved from yeast to human. We hope that results from our yeast prion studies will provide valuable information on the complex etiology of the devastating prion diseases.
Our laboratory is also interested in investigating how common the prion phenomenon in biology is. We wish to identify potential prion proteins from yeast and other non-yeast model organisms through a combined approach of bioinformatics and genetic screenings. Our ultimate goal is to uncover the mechanisms governing the prion conformational switch and to understand the biological significance of the protein conformation based prion-like inheritance.
For more information, visit the faculty profile of Liming Li, PhD.
Publications
See Dr. Li's publications in PubMed.
Contact

Xiao-Nan Li
Professor, Pediatrics (Hematology, Oncology, and Stem Cell Transplantation)
Understanding tumor biology and implementing preclinical drug testing of malignant brain tumors

Xiao-Nan Li
Research Area(s): Cancer Biology; Neurobiology; Pharmacology
Our research work focuses on molecular neuro-oncology and experimental therapeutics of malignant brain tumors. Our goal is to develop more effective and less toxic therapies for children with malignant brain tumors to significantly improve the clinical outcomes. 1) Clinically relevant and molecularly accurate animal models: We have optimized a protocol for to implant a patient’s tumor cells directly into the matched location in mouse brains. Our laboratory has developed >130 patient derived orthotopic xenograft (PDOX or orthotopic PDX) models of pediatric brain tumors, including high-grade glioma/glioblastoma (GBM), medulloblastoma (MB), ependymoma (EPN), diffuse intrinsic pontine glioma (DIPG), atypical teratoid/rhabdoid tumor (ATRT), Embryonal tumors with multilayered rosettes (ETMR), CNS germinoma, and pleomorphic xanthoastrocytoma (PXA), as well as ~20 models of adult GBM and meningioma. 2) All the models are subjected to detailed histopathological and comprehensive molecular characterizations through global gene expression (RNAseq, single cell RNAseq), whole genome sequencing, DNA copy number analysis, whole genome DNA methylation analysis during serial in vivo subtransplantations to confirm thee cellular and molecular fidelities of the patient-specific PDOX models. 3) Recognizing the need of in vitro model system, we are also utilizing our PDOX model system to develop long term cultures/cell lines as monolayer, 3D neurospheres and organoids. 4) Using this unique in vitro and in vivo model system, we are actively engaged in the understanding of tumor biology and the testing of new therapeutic strategies.
Current Projects
- Understanding the mechanisms of brain tumor invasion of pediatric GBM
- Discovery of cellular and molecular drivers of medulloblastoma metastasis
- Investigating the regulation of cell cycle progression, particularly from quiescent G0 to active G1 phase
- Implementing unbiased high-throughput combinatory drug screening against highly malignant brain tumors using a novel drug library composed of 17,000 drugs and investigational agents
- Identifying cellular origins and molecular drivers of therapy resistance and tumor relapse
- Evaluating therapeutic efficacy and elucidation of mechanisms of action of novel anti-cancer therapies in vivo in PDOX models for the initiation of clinical trials.
For more information, visit the faculty profile of Xiao-Nan Li, MD, PhD.
Publications
See Dr. Li's publications.
Contact

Huiping Liu
Associate Professor, Pharmacology, Medicine (Hematology and Oncology)
Targeting cancer stem cells and exosomes in metastasis using cutting-edge technology and novel therapeutics

Huiping Liu
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Immunology
The Liu lab studies the molecular and cellular mechanisms underlying cancer stem cells (CSCs) and metastasis through four ongoing interactive basic and translational research projects: (1) to understand CSCs, circulating tumor cells (CTCs) and their interactions with immune cells in metastasis; (2) to dissect the role of secreted and circulating exosomes in CSC functions; (3) to target CSCs with novel therapeutics, exosomes and nanoparticles in combination with immunotherapy; (4) to develop CTC and circulating exosome-based biomarkers for cancer diagnosis, therapy response and predictive prognosis.
Recent Findings
- CSCs seed metastases of breast cancer.
- A rapid, automated surface protein profiling of single circulating exosomes in human blood. (PMID: 27819324)
- Micro-206 inhibits stemness and metastasis of breast cancer by targeting MKL1/IL11 pathway. (PMID: 27435395)
- Differentiation and loss of malignant character of spontaneous pulmonary metastases in patient-derived breast cancer models. (PMID: 25339353)
- MicroRNA-30c inhibits human breast tumor chemotherapy resistance by regulating TWF1 and IL-11.(PMID: 23340433)
Current Projects
- Identify how CSCs crosstalk between themselves and interplay with immune cells in circulation thus developing innovative anti-CSC targeting therapeutics.
- Single cell RNA sequencing of CSCs and metastatic tumors.
- Dissect the role of exosomes in CSC functions and regulation of exosome functions by CSCs.
- Develop CSC-targeting novel microRNA therapeutics and exosome delivery tools.
- Discover CTC and circulating exosome-based clinical biomarkers for cancer diagnosis, treatment monitoring and prognosis.
For lab information and more, see Dr. Liu's faculty profile.
Publications
See Dr. Liu's publications on PubMed.
Contact
Contact Dr. Liu at 312-503-5248.

Yan Liu
Associate Professor, Medicine (Hematology and Oncology)
Hematopoietic stem cell self-renewal and pathogenesis of myeloid malignancies

Yan Liu
Research Area(s): Cancer Biology; Developmental and Stem Cell Biology
The Liu laboratory is interested in investigating the molecular mechanisms governing normal and malignant hematopoiesis, with an emphasis on understanding hematopoietic stem cell (HSC) self-renewal and pathogenesis of myeloid malignancies, including myelodysplastic syndromes (MDS) and acute myeloid leukemia (AML). Our long-term goals are to identify novel regulators of HSC self-renewal, understand the molecular mechanisms regulating their function, and develop novel therapeutic strategies to eliminate leukemia stem cells (LSCs) and improve leukemia treatment. We utilize molecular, genetic, immunological, biochemical, and pharmacological approaches as well as unbiased genome wide studies, including RNA-seq, ChIP-seq, ATAC-seq, and cytokine arrays, to investigate the molecular basis of HSC self-renewal and leukemogenesis. Major areas of focus include: 1) The role of tumor suppressor p53 in CHIP progression and pathogenesis of MDS; 2) Phosphatase PRL2 in HSC self-renewal and leukemogenesis; 3) Polycomb Repressive Complex 1 (PRC1) in HSC self-renewal and terminal differentiation.
For more information, visit Dr. Liu's faculty profile page.
Publications
View Dr. Liu's publications at PubMed.
Contact

Yaping Liu
Assistant Professor, Biochemistry and Molecular Genetics, Neurology - Ken and Ruth Davee Department
Epigenomics and bioinformatics in single-cell and liquid biopsy to dissect the function of non-coding genomes in human diseases

Yaping Liu
Research Area(s): Cancer Biology; Cell Biology; Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Microbiology; Neurobiology
The Liu lab is interested in understanding the gene regulatory roles of non-coding genetic variants and therefore bridging the gaps between genetic variations and phenotypic variations at different physiological and pathological conditions, such as cancers, developmental and neurodegenerative disorders. To achieve these goals, (1) we are developing high-throughput single-molecule and single-cell multi-omics experimental approaches to simultaneously capture multiple epigenomes, such as DNA methylation and 3D genome, from the same DNA molecules and single cells; (2) we are also developing computational methods to non-invasively monitor the in vivo epigenomes by the fragmentation patterns of circulating cell-free DNA and therefore enabling the detection of early-stage diseases (e.g. cancers) and prediction of the cancer patients' responses to the immunotherapy.
For more information, visit Dr. Liu's faculty profile page.
Publications
View Dr. Liu's publications at PubMed.
Contact

Richard M Longnecker
Professor, Microbiology-Immunology
Epstein-Barr virus (EBV) and herpes simplex virus (HSV) entry, replication and pathogenesis

Richard M Longnecker
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Immunology; Microbiology; Neurobiology; Pharmacology
Research in the Longnecker laboratory focuses on herpes simplex virus (HSV) and Epstein-Barr virus (EBV). These viruses typically cause self-limiting disease within the human population but both can be associated with serious complications. EBV is associated with variety of hematopoietic cancers such as African Burkitt lymphoma, Hodgkin Lymphoma and adult T-cell leukemia. EBV-associated lymphoproliferative disease occurs in individuals with congenital or acquired cellular immune deficiencies. The two notable epithelial diseases associated with EBV infection are nasopharyngeal cancer and oral hairy leukoplakia. Similar to EBV, HSV latent infections are very common in humans. HSV typically does not cause severe disease but is associated with localized mucocutaneous lesions, but in some cases can cause meningitis and encephalitis. The Longnecker laboratory focuses on several aspects of EBV and HSV replication and pathogenesis. First, the molecular basis EBV transformation and how it relates to cancer is being investigated. The laboratory is currently screening selective inhibitors that may be beneficial in EBV-associated cancers such as Hodgkin lymphoma, Burkitt lymphoma and proliferative disorders that occur in HIV/AIDS and transplant patients. Second, the laboratory is investigating herpesvirus latency in the human host and pathogenesis associated with infections in humans. In this regard, the laboratory is developing animal models for EBV and HSV infections. Finally, the laboratory is investigating the function of herpesvirus encoded proteins and the cellular receptors that are important for infection both using in vivo culture models as well as animal models. Ultimately, studies by the Longnecker laboratory may provide insight for the development of novel therapeutics for the treatment of herpesvirus infections in humans and better understanding of the herpesvirus life cycle in the human host
For lab information and more, see Dr. Longnecker's faculty profile.
Publications
See Dr. Longnecker's publications on PubMed.
Contact
Contact Dr. Longnecker at 312-503-0467 or the lab at 312-503-0468 or 312-503-9783.

Ramon Lorenzo Redondo
Assistant Professor, Medicine (Infectious Diseases)
Viral evolution and virus-host interaction to develop better prevention and cure strategies against viral pathogens

Ramon Lorenzo Redondo
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Microbiology
As a researcher in the Division of Infectious Diseases in Northwestern University, my work combines virology and evolutionary biology to study viral evolution and the interaction between viruses and the host during infection. My main interests are RNA viruses, molecular evolution, and genomics. The ultimate goal of my research is to understand the virus-host system and its evolutionary properties in order to develop the best treatments and prevention strategies for human viral infections. During my career, I have led published studies on the evolution of HIV-1 and contributed greatly to the field of RNA virus host-virus genetics. I utilize state-of-the-art sequencing technologies and a combination of evolutionary biology, bioinformatics, statistical modeling, complex data analysis, and phylogenetics to answer fundamental questions about viral population dynamics and viral interaction with the host. My lab has adapted and developed viral deep sequencing pipelines and host genomics techniques, including bulk and single cell transcriptomics, as well as innovative spatial transcriptomics pipelines, to study the virus-host system. I am the director of bioinformatics of the Northwestern University Center for Pathogen Genomics and Microbial Evolution (CPGME) and a faculty member of Northwestern’s Institute of Global Health, the Third Coast Center for AIDS Research (TC CFAR), and the Buehler Center for Health Policy and Economics.
My research projects expand from clinically relevant studies that examine viral infection dynamics at the human population level both in national and global health settings, to studies that intend to comprehend viral reservoirs, with special focus on the HIV-1 cure field, as well as intra-host viral evolution and virus-host interaction.
For lab information and more, see Dr. Lorenzo-Redondo's faculty profile.
Publications
See Dr. Lorenzo-Redondo's publications on PubMed.
Contact
Contact Dr. Lorenzo-Redondo

William L Lowe, Jr.
Professor Emeritus, Medicine (Endocrinology, Metabolism and Molecular Medicine)
Genetic determinants of maternal metabolism and fetal growth

William L Lowe, Jr.
Research Area(s): Cardiovascular and Renal Biology; Genetics, Epigenetics and Genomics
A major interest of the Lowe laboratory is genetic determinants of maternal metabolism during pregnancy and the interaction between the intrauterine environment and genetics in determining size at birth. This interest is being addressed using DNA and phenotype information from ~16,000 mothers and their babies who participated in the Hyperglycemia and Adverse Pregnancy Outcome (HAPO) Study. A genome wide association study using DNA from mothers and babies from four different ancestry groups has been performed, with several different loci demonstrating genome-wide significant association with maternal and fetal traits. Replication studies have confirmed the identified associations. Studies are now underway now to identify the causal variants and their functional impact. In related studies performed with investigators at Duke, targeted and untargeted metabolomic studies are underway to determine whether metabolic signatures characteristic of maternal obesity and/or hyperglycemia can be identified in mothers and babies. Integration of metabolomic and genomic data is also planned to more fully characterize maternal metabolism during pregnancy and its interaction with fetal growth. Finally, a HAPO Follow-Up Study has been initiated in which a subset of the HAPO mothers and babies (now 8-12 years of age) will be recruited to examine the hypothesis that maternal glucose levels during pregnancy are positively correlated with metabolic measures in childhood, including adiposity, lipidemia, glycemia and blood pressure.
For further information visit Dr. Lowe's faculty profile page
Publications
View Dr. Lowe's publications at PubMed
Contact
Phone 312-503-2539

Yuan Luo
Professor, Preventive Medicine (Biostatistics and Informatics), Pediatrics
Professor, McCormick School of Engineering
Medical and clinical applications of machine learning, natural language processing, time series analysis, integrative genomic analysis and big data analytics

Yuan Luo
Research Area(s): Cardiovascular and Renal Biology; Computational Biology and Bioinformatics
For more information visit Dr. Luo's faculty profile page
Publications
See Dr. Luo's publications on PubMed.
Contact
Phone 312-908-7914

Yongchao C Ma
Professor, Pediatrics
Regulation of Motor Neuron and Dopaminergic Neuron Function in Development and Disease

Yongchao C Ma
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Neurobiology; Pharmacology
Regulation of Neural Stem Cell Development and Neurodegeneration by RNA Methylation
RNA methylation on N6-adenosine (m6A) has emerged as a crucial regulator of RNA function and metabolism, directly influencing protein expression and biological functions. We recently identified novel m6A RNA readers that interpret RNA methylation to control diverse aspects of RNA biology, including its localization, degradation and translation. We are investigating how RNA methylation regulates fundamental processes in neural development and degeneration, such as neurogenesis, neural stem cell differentiation, and neuroinflammation. We also explore its role in the pathogenesis of autism, ALS and other neurological disorders.
To uncover these molecular mechanisms, we employ a combination of genetic, biochemical and cell biological approaches, using genetically modified mice and human induced pluripotent stem (iPS) cells. We employ next generation sequencing and data analysis to map RNA methylation sites, and we develop novel RNA methylation-based therapeutics through high-throughput drug screening.
Regulation of Mitochondrial Function in Motor Neuron Degeneration
Spinal muscular atrophy (SMA), the leading genetic cause of infant mortality, affects one in every eight thousand live births, while amyotrophic lateral sclerosis (ALS), also known as Lou Gehrig’s disease, is the most common motor neuron disease in adults. Both conditions are characterized by the selective degeneration of spinal motor neurons. Our group focuses on understanding the mechanisms that regulate motor neuron development and function, as well as the specific vulnerability of motor neurons in SMA and ALS. To address these questions, we perform single-cell RNA sequencing and signaling pathway analysis, using SMA and ALS mouse models along with stem cell-derived neurons. A key focus of our research is the non-canonical role of mitochondria as signaling hubs, regulating epigenetic and metabolic changes that contribute to neurodegeneration in SMA and ALS.
Aging and Mitochondrial Oxidant Defense in Dopaminergic Neuron Degeneration of Parkinson’s Disease
Dopaminergic neurons in the ventral midbrain control movement, emotional behavior and reward mechanisms. Dysfunction of these neurons is implicated in Parkinson’s disease (PD), drug addiction, depression and schizophrenia. Our group studies the genetic and epigenetic mechanisms regulating dopaminergic neuron functions in disease and aging conditions. We are particularly interested in how aging and impaired mitochondrial oxidative defense contribute to the selective degeneration of dopaminergic neurons in PD through transcriptional and epigenetic changes. For these studies, we use mouse models, cultured neurons, and iPS cells to explore the underlying molecular mechanisms.
For more information visit Dr. Ma's faculty profile and Dr. Ma's lab website
Publications
View Dr. Ma's publications at PubMed
Contact
Phone 312-503-7338.

Kyle L MacQuarrie
Assistant Professor, Pediatrics (Hematology, Oncology, and Stem Cell Transplantation)
Intersection of developmental and cancer biology in pediatric solid tumors

Kyle L MacQuarrie
Research Area(s): Cancer Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics
Pediatric cancer long-term survival rates overall have climbed steadily for multiple decades. However, there are still multiple types of pediatric tumors that have seen little improvement in survival rates during this time, including the most common pediatric sarcomas. The MacQuarrie lab focuses on the intersection of normal developmental biology and cancer biology in these sorts of tumors, with the aim of identifying both normal developmental events that are absent in the tumors and tumor-specific events that are absent in normal development and then leveraging these differences to improve clinical outcomes in children long-term.
Our current studies focus on nuclear and genomic organization, and particularly the geometric organization of chromatin and genes in nuclear space. We most commonly study rhabdomyosarcoma, a pediatric tumor of skeletal muscle, as well as normal developing skeletal muscle cells. Using cell culture models and primary tumor samples, we use ‘classic’ molecular and cellular biology techniques, a variety of next-generation sequencing techniques, and microscopy (including super-resolution approaches) to investigate our scientific hypotheses.
For more information view Dr. MacQuarrie's Faculty Profile
Publications
View Dr. MacQuarrie's publications by PubMed
Contact
Michelle L Macy
Professor, Pediatrics (Emergency Medicine)
Advancing equity and precision prevention in childhood by leveraging data and digital health tools
Michelle L Macy
Research Area(s): Computational Biology and Bioinformatics
Dr. Macy is a pediatric emergency physician and NIH-funded clinician scientist who has focused on addressing leading causes of unintentional injury in childhood, motor vehicle collision and drowning. She develops and studies new approaches to promote child passenger safety best practices through tailored health communications, digital health tools, and motivational interviewing. She collaborates with community leaders in metropolitan Chicago to capture data on drowning events and evaluate community-based interventions to teach water safety to groups who historically have been excluded from learning to swim. With the Injury Equity Framework (Kendi & Macy, NEJM 2023), she is leading a shift in the injury prevention field away from individual behavior toward systematic assessments of historic and current contributors and countermeasures to inequitable outcomes within the context of individual, family, community, and societal factors. Dr. Macy serves Scientific Director for Community, Population Health and Outcomes in Stanley Manne Children’s Research Institute at Ann & Robert H. Lurie Children’s Hospital of Chicago. She is a co-investigator in the Measurement Core for the Environmental Influences on Child Health Outcomes (ECHO) Program and for the Mental Health Earlier ALACRITY Center within the Institute for Innovations in Developmental Sciences. She also contributes to data analysis and interpretation for Pediatric and Emergency Medicine Value Based Care initiatives in Illinois and Michigan.
For more information visit Dr. Macy's faculty profile
Publications
View Dr. Macy's publications at PubMed
Contact

Aline Martin
Associate Professor, Medicine (Nephrology and Hypertension)
Role of the skeleton in the endocrine regulation of mineral metabolism and the cardiovascular complications of mineral and bone diseases

Aline Martin
Research Area(s): Cardiovascular and Renal Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology
Our research program focuses on the contribution of the skeleton to the mineral balance in the body. Bone produces a hormone, Fibroblast Growth Factor (FGF)-23, that participates in this balance. However in mineral metabolism disorders, such as in chronic kidney disease, the massive production of FGF23 is associated with negative outcomes and mortality. By understanding the mechanisms that control the production of FGF23, our goal is to develop new therapeutic strategies and improve outcomes in mineral metabolism disorders. To this goal, we perform basic and translational research using a combination of genetics, molecular biology, proteomics, histology and advanced imaging techniques.
A major focus of the lab is to investigate the transcriptional and post-translational regulation of FGF23 within the bone cells. In particular, we study the specific role of a known regulator of FGF23, Dentin Matrix Protein 1 (DMP1), on these regulations and on osteocyte biology in the context of diseases associated with FGF23 excess (chronic kidney disease, hypophosphatemic rickets …). A second focus is to investigate the mechanisms involved in negative outcomes associated with FGF23 excess, including bone mineralization defects, cardiac hypertrophy and cognitive defects. Our team works in collaboration with the Center for Translational Metabolism and Health and the Division of Cardiology at Northwestern, and with multiple additional collaborators and partnerships around the world.
The Martin Lab is sponsored by the National Institute of Health, National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) and by the Northwestern Women’s Health Research Institute.
Publications
For more information view Dr. Martin's Faculty Profile or view publications by PubMed
Contact Us
Contact Dr. Martin at 312-503-4160 or the Martin Lab at 312-503-4805, or by email.

Elena Martinelli
Professor, Medicine (Infectious Diseases), Microbiology-Immunology
Mucosal Immunity and HIV host-viral interactions

Elena Martinelli
Research Area(s): Immunology; Microbiology; Pharmacology; Reproductive Biology
Our research focuses on understanding immunity in mucosal tissues such as the gastrointestinal tract and the female reproductive tract and how the balance of tolerogenic and inflammatory responses are maintained during a persistent viral infection such as HIV. At the moment, the research in the lab is particularly focused on dissecting HIV persistence during antiretroviral therapy and how it is modulated by stimuli present in the tissue microenvironment (including the role of co-pathogens, hormones, cytokines, metabolites etc).
HIV persistence represents a major obstacle to HIV cure and a major focus on the laboratory is testing novel therapeutic strategies that can help developing a functional cure for HIV.
We use a variety of experimental models that include in vitro, ex vivo and in vivo systems ranging from cell lines, primary cell models, tissue explants, gut organoids, mice and non-human primates (NHP). We have active collaborations with primate centers in Louisiana where our NHP work is being conducted and most of the lab efforts center on NHP studies and samples and tissues collected from those studies. We apply and help developing novel imaging techniques such as immunoPET/CT to visualize virus and immune system components in NHP and conduct PET-guided sampling in NHP models. Single cell, spatial biology, and omics techniques (including transcriptomics, proteomics, and metabolomics) are applied on the tissue site/s of interest (an in in vitro studies) to dissect changes in viral-host interactions at a molecular and spatial level.
Publications
For more information view Dr. Martinelli's Faculty Profile or view publications by PubMed
Contact

Daniela E Matei
Adjunct Lecturer, Obstetrics and Gynecology
Mechanisms of ovarian cancer metastasis and novel therapeutics for ovarian cancer

Daniela E Matei
Research Area(s): Cancer Biology; Pharmacology; Reproductive Biology
My laboratory studies mechanisms of ovarian cancer metastasis and novel therapeutics for ovarian cancer. The general theme is translation between bench and clinic; with laboratory research forming the foundation for clinical experiments.
One direction of investigation relates to the interaction between cancer cells and the peritoneal stroma. We described the functions of tissue transglutaminase as an interacting partner of b-integrins and regulator of peritoneal metastasis. Based on new mechanistic insight into the roles of this enzyme in ovarian cancer, we discovered and began characterizing new small molecule inhibitors for the transglutaminase-fibronectin-integrin interaction that are being developed as anti-cancer agents. We are studying these new inhibitors in-vitro and in in-vivo models of ovarian cancer metastasis.
Another area of research focusses on the characteristics and therapeutic vulnerabilities of ovarian cancer stem cells. We used small molecule inhibitors that target ALDH1A1 to block the tumorigenic capacity of these cancer-initiating cells. We are studying how ALDH1A1 inhibitors alter stem cell specific signaling and how ALDH1A1 is involved in maintaining the cancer stem cell properties.
More recently we identified new metabolic alterations involving fatty acids desaturation in cancer stem cells. We have targeted lipid metabolism using small molecule inhibitors and are studying the mechanisms by which these metabolic changes contribute to the maintenance and tumorigenicity of cancer stem cells. Future goals are to refine the use of ALDH and fatty acid desaturases inhibitors to target cancer stem cells residual after chemotherapy and to eradicate the disease.
Another important direction of investigation is epigenetic modulation as a method of resensitization to platinum in ovarian cancer. We successfully brought to the clinic the concept that epigenetic modulation re-sensitizes chemotherapy-resistant ovarian tumors to carboplatin. I led a randomized multi-institutional clinical trial testing the effects of DNA hypomethylating agents and carboplatin compared to standard chemotherapy. We are now analyzing the genome and epigenome of platinum resistant ovarian cancer using specimens from this trial. We have identified several pathways that are associated with platinum resistance and respond to hypomethylating agents. We have designed a new strategy to target pathway-specific DNA methylation and are testing the effects of this intervention on cell signaling and gene expression profiles in ovarian cancer cells.
Publications
View Dr. Matei's publications on PubMed
Contact
Phone 312 503-4853

Elizabeth M McNally
Professor, Medicine (Cardiology), Biochemistry and Molecular Genetics
Genetic mechanisms responsible for inherited human diseases

Elizabeth M McNally
Research Area(s): Cardiovascular and Renal Biology; Genetics, Epigenetics and Genomics
My laboratory studies genetic mechanisms responsible for inherited human diseases including heart failure, cardiomyopathy, muscular dystrophy, arrhythmias, aortic aneurysms. Working with individuals and families, we are defining the genetic mutations that cause these disorders. By establishing models for these disorders, we can now begin to develop and test new therapies, including genetic correction and gene editing.
For lab information and more, see Dr. McNally's faculty profile or visit the McNally Laboratory site.
Publications
See Dr. McNally's publications on PubMed.
Contact
Email Dr. McNally
Phone 312-503-5600
Joshua J Meeks
Associate Professor, Urology, Biochemistry and Molecular Genetics
Genetic and epigenetic changes in bladder cancer, and immuno-oncology in bladder cancer
Joshua J Meeks
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Immunology
The Meeks lab is investigating the epigenetics and genetic mutations associated with cancer biology. Specifically, he is studying how chromatin remodeling genes play a role in bladder cancer. In addition, he is investigating the “driver mutations found in bladder cancer. In the future, he hopes to develop novel systemic and intravesical therapies to improve survival of patients with bladder cancer.
In the United States, there are an estimated 72,570 new cases of bladder cancer each year. Dr. Meeks is conducting innovative research to increase our understanding of the biology of bladder cancer and to identify new therapies and technologies for bladder cancer in order to improve quality of life for our patients. In this podcast, Joshua Meeks, MD, PhD, shares how his team of scientists are involved in three active trials focused on genetic and epigenetic changes in bladder cancer, as well as immuno-oncology in bladder cancer. Listen here >>
Dr. Meeks is investigating the gender disparities in bladder cancer by dissecting the tumor and immune mechanisms of resistance to chemotherapy and immunotherapy. This research may translate into novel pathways and potential therapeutic targets to improve outcomes and reduce gender disparities in bladder cancer. In this video, Meeks shares details about his work. Watch here >>
For more information visit Dr. Meeks' faculty profile or the Meeks Lab website
Publications
Please see PubMed for a full list of publications.
Contact

Johanna Melo-Cardenas
Assistant Professor, Pathology
Inflammatory signaling in normal and malignant blood development

Johanna Melo-Cardenas
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology; Immunology; Pharmacology
We study the function of inflammatory regulators and their impact on hematopoietic cells (normal and malignant). By leveraging advanced technologies, mouse models and human samples, we aim to discover new strategies to slow or stop the progression of blood cancers.
For more information visit Dr. Melo-Cardenas' faculty profile
Publications
Please see PubMed for a full list of publications.
Contact

Marc L Mendillo
Associate Professor, Biochemistry and Molecular Genetics
Cellular stress response systems in malignancies

Marc L Mendillo
Research Area(s): Cancer Biology; Cell Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Pharmacology
The cellular stress response systems guard the proteome from diverse endogenous and environmental insults to maintain the fitness of the organism. Ironically, this pro-survival system can act to the detriment of the host to enable tumor cells accommodate to the myriad stresses associated with malignancy. Our long-term goals are to identify and characterize the systems that promote protein homeostasis, understand how these systems are co-opted and perturbed in malignancy, and ultimately identify means to manipulate them for therapeutic benefit. To accomplish these goals our group bridges biochemical, genetic and chemical biology approaches with systematic high-throughput and genomic methods.
For lab information, publications and more, see Dr. Mendillo’s faculty profile.
Publications
View Dr. Mendillo's publications at PubMed
Contact
Phone 312-503-5685

Richard J Miller
Professor Emeritus, Pharmacology
Molecular aspects of nerve cell communication and neurodegenerative disease

Richard J Miller
Research Area(s): Cell Biology; Neurobiology; Pharmacology
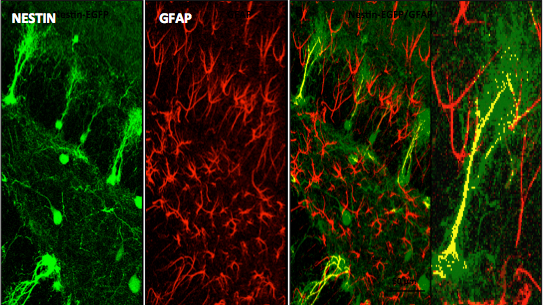
Co-localization of Nestin and GFAP in the DG of Nestin-EGFP Transgenic Reporter Mice
The laboratory led by Richard Miller, PhD, is interested in studying molecular aspects of nerve cell communication. One of our major interests has been to understand the structure and function of calcium channels. The influx of Ca into neurons through these channels is important for many reasons, including the release of neurotransmitters. We have identified a family of molecules that act as Ca channels in neurons and other types of cells. Each of these molecules has slightly different properties that underlie different neuronal functions. We have analyzed the properties of these molecules by examining their electrophysiological properties following their expression in heterologous expression systems and imaging techniques. Furthermore, we have generated calcium channel knockout mice that have interesting properties such as altered pain thresholds, seizures and memory deficits. We have also been interested in how Ca channels can be regulated by the activation of Gprotein coupled receptors. We have been analyzing the interaction of Gprotein subunits with Ca channels using FRET imaging and other techniques.
Other projects in our laboratory are aim to understanding the molecular basis of neurodegenerative disease. We study Alzheimer's disease, Amyotrophic lateral sclerosis (Lou Gehrig's disease), HIV-1 related dementia and other neuropathological conditions. In the case of HIV-1 infection, we have been examining the properties and functions of HIV-1 receptors on neurons. These receptors are known to be receptors for chemokines -small proteins that are known to direct the functions of the immune system. We have shown that neurons express many types of chemokine receptors and that activation of these receptors can produce both short and long term effects on neurons. Activation of chemokine receptors expressed by sensory neurons produces neuronal excitation and pain. Activation of chemokine receptors on hippocampal neurons has a prosurvival effect, whereas binding of HIV-1 to these receptors induces apoptosis. We are studying the molecular mechanisms that produce this diverse effects with a view to understanding the molecular basis for HIV-1 related dementias.
For lab information and more, see Dr. Miller’s faculty profile.
See Dr. Miller’s blog “The Keys to all Mythologies: Science, Medicine and Magic” to read articles concerning scientific topics of current interest as well as historical accounts of scientific issues.
Publications
See Dr. Miller's publications on PubMed.
Contact
Contact Dr. Miller at 312-503-3211.

Stephen D Miller
Professor Emeritus, Microbiology-Immunology
Mechanisms of pathogenesis and immune regulation of autoimmune disease, allergy and tissue/organ transplantation

Stephen D Miller
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Immunology; Pharmacology
The laboratory is interested in understanding the mechanisms underlying the pathogenesis and immunoregulation of T cell-mediated autoimmune diseases, allergic disease and rejection of tissue and organ transplants. In particular, we are studying the therapeutic use of short-term administration of costimulatory molecule agonists/antagonists and specific immune tolerance induced by infusion of antigen-coupled apoptotic cells and PLG nanoparticles for the treatment of animal models of multiple sclerosis and type 1 diabetes, allergic airway disease, as well as using tolerance for specific prevention of rejection of allogeneic and xenogeneic tissue and organ transplants.
For lab information and more, see Dr. Miller's faculty profile.
Publications
See Dr. Miller's publications on PubMed.
Contact
Contact Dr. Miller at 312-503-7674 or the lab at 312-503-1449.

Booki Min
Professor, Microbiology-Immunology
Regulatory T cells in inflammation

Booki Min
Research Area(s): Cancer Biology; Immunology
The immune system is tightly controlled by multiple mechanisms, and Foxp3+ regulatory T (Treg) cells are prominent active regulators of immunity and tolerance. Defects in Treg cell generation or function result in uncontrolled systemic autoimmune inflammation. Despite extensive investigation in Treg cell biology, our understanding the mechanisms underlying Treg cell development and functions still remains incomplete. There is increasing evidence that Treg cell functions can be compromised under certain conditions, and such dysregulation is thought to be a contributing factor of chronic inflammatory conditions. Our laboratory studies both cellular and molecular factors that control Treg cell functions.
There are three major research projects currently underway in the laboratory.
1. IL-27 and Treg cells: Earlier studies had identified that IL-27, an immune regulatory cytokine produced by activated APCs, plays a non-redundant role in regulating Treg cell function. IL-27 acts on the IL-27 specific receptors (made of IL-27Ra and gp130 subunit) expressed on multiple cell types, primarily lymphocytes. Using various genetic approaches, our research focuses on identifying key source of IL-27 in autoimmune inflammation in the central nervous system and underlying mechanism by which IL-27 controls Treg cell function.
2. Glucocorticoids, miR-342, and Treg cells: We recently reported a novel role of Treg cells during glucocorticoid-induced treatment of chronic inflammation. In this study, we discovered a novel micro-RNA-342 molecule that is induced by steroid treatment in Treg cells and directly controls Treg cell metabolism. We are investigating the underlying mechanism by which this new micro-RNA-342 controls Treg cell function.
3. Lag3 and Treg cells: Lag3 is a new immune checkpoint molecule implicated in negatively regulating T cell functions. We discovered that Lag3 is induced by IL-27 stimulation in Treg cells and that Lag3 expression in Treg cells is critical for their suppressive function. We have generated several new mouse models in which Lag3 function and signaling pathways are targeted in a cell type specific manner. These novel animal models will allow us to dissect underlying mechanism of Lag3 in Treg cells and to identify potential therapeutic strategies not only to inhibit inflammation but also to enhance anti-tumor immunity by targeting Lag3 in Treg cells.
For lab information and more, see Dr. Min’s faculty profile and laboratory website.
Publications
See Dr. Min's publications.
Contact
Contact Dr. Min at 312-503-1805.

Alexander V Misharin
Associate Professor, Medicine (Pulmonary and Critical Care)
Lung repair after injury and infection

Alexander V Misharin
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Immunology; Metabolism and Endocrinology; Neurobiology
The Misharin laboratory studies mechanisms governing normal and aberrant lung repair after injury, with the ultimate goal of making our findings translatable and relevant for clinical practice and human health. Specifically, we are focusing on interactions between the immune, epithelial and stromal lung cells in the context of influenza A virus-induced and SARS-CoV-2-induced pneumonia (McQuattie et al., JCI, 2020; Bharat et al., Sci Trans Med, 2020; Grant et al., Nature, 2021), pulmonary fibrosis (Reyfman et al., AJRCCM, 2019; Joshi et al., ERJ, 2020; Watanabe et al., PNAS, 2021), chronic lung allograft dysfunction, and aging. Our research also covers extra-pulmonary complications of viral pneumonia, such as muscle dysfunction and cognitive impairment (Runyan et al., Aging Cell, 2020). While we recognize that multiple cell types contribute to lung repair after the injury and act in tight orchestration with each other, we are dissecting these complex interactions by focusing our research on macrophages – a cell type that is ubiquitously present in every organ and tissue. Macrophages actively contribute to injury and repair. They also evolved to adapt to the surrounding environment and thus can serve as an accurate “mirror”, which reflects the state of the tissue during the steady-state, injury, and repair (see our review: Watanabe et al., JCI, 2019). We use biosamples and clinical data from patients with lung diseases and combine them with modern single cell genomic and machine learning approaches to drive our discoveries (Lotfollahi et al., Nat Biotech, 2022; Gao et al., JCI, 2023; Sikkema et al., Nat Med, 2023), which we then validate in causal animal models. We actively partner with industry to bring our discoveries back to bedside.
For lab information and more, see Dr. Misharin’s faculty profile
Publications
See Dr. Misharin's publications.
Contact
Contact Dr. Misharin

Jason Miska
Assistant Professor, Neurological Surgery
Metabolism of immune cells in brain tumors

Jason Miska
Research Area(s): Cancer Biology; Immunology; Metabolism and Endocrinology; Neurobiology
The goal of the Miska laboratory's is to determine how the metabolism of immune cells within brain tumors contributes to immune suppression and tumor recurrence. Furthermore, we seek to manipulate these metabolic pathways in a clinically relevant manner to improve patient outcomes for this deadly disease. Currently, we are exploring how the unique metabolism of tumor-associated myeloid cells (TAMCs) promote their survival, immunosuppression, and tumor brain progression. We have discovered that inhibiting the downstream products of arginine metabolism is a useful strategy for promoting anti-tumor immune responses. Our laboratory also performs immunological monitoring for clinical trials in brain tumor patients by monitoring immune phenotypes, T-cell reactivity, and changes in systemic cytokines that occur with therapeutic administration.
For lab information and more, see Dr. Miska’s faculty profile
Publications
See Dr. Miska's publications.
Contact
Contact Dr. Miska

Brian J Mitchell
Associate Professor, Cell and Developmental Biology
Integration of signaling and cytoskeletal dynamics on diverse developmental processes including centriole amplification, cell migration and cell polarity

Brian J Mitchell
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology
Centrioles are microtubule based structures with nine fold symmetry that are involved in both centrosome organization and aster formation during cell division. During the normal cell cycle centrioles duplicate once, generating a mother/daughter pair and in most post-mitotic vertebrate cells the mother centriole then goes on to form the basal body of a sensory cilium. Abnormalities in the duplication of centrioles (and centrosomes) are prevalent in many cancers suggesting a link between centriole duplication and cancer progression. We study what factors limit centriole duplication from a novel direction with the use of Xenopus motile ciliated cells. Ciliated cells are unique among vertebrate cells in that they generate hundreds of centrioles (basal bodies) therefore providing a great system for studying the regulation of centriole duplication. Understanding how nature has overcome the typically tight regulation of centriole duplication will lend insight into the molecular mechanisms of cancer progression.
Tissue development and homeostasis requires dramatic remodeling as new cells migrate into an epithelium. How migrating cells breakdown junctional barriers during development or during diseases processes such as metastasis is poorly understood at the molecular level. During the early development of Xenopus embryos, distinct cell types join the outer epithelium in a process called radial intercalation. We are interested in the molecular mechanisms that regulate both the migration of these cells as well as the tissue remodeling that occurs to accommodate them.
The ability of ciliated epithelia to generate directed fluid flow is an important aspect of diverse developmental and physiological processes including proper respiratory function. To achieve directed flow, ciliated cells must generate 100-200 cilia that are polarized along a common axis both within and between cells. My lab is currently working towards understanding the molecular mechanisms for how cell polarity is coordinated as well as how individual cilia interpret the cells polarity. We have determined that ciliated cells receive polarity cues via the non-canonical Wnt/Planar Cell Polarity (PCP) pathway, but the details of this are still poorly understood. Additionally, the PCP pathway is known to influence a cells cytoskeleton dynamics and a main goal is to understand how this influences the ability of individual cilia to coordinate their polarity.
For lab information and more, see Dr. Mitchell's faculty profile and lab website.
Publications
See Dr. Mitchell's publications on PubMed.
Contact
Contact Dr. Mitchell at 312-503-9251.

David C Mohr
Professor, Preventive Medicine (Behavioral Medicine), Medical Social Sciences (Intervention Science), Psychiatry and Behavioral Sciences
Design, developing, evaluating, and implementing technology-assisted behavioral and psychological interventions

David C Mohr
Research Area(s): Computational Biology and Bioinformatics
David C. Mohr, PhD, is the Director of the Northwestern University Center for Behavioral Intervention Technologies (CBITs). Dr. Mohr’s expertise is in the design, development, evaluation, and implementation of technology-assisted behavioral and psychological interventions. These technologies use mobile phones, tablets, computers, and sensors to support patient behaviors related to health, mental health, and wellness. In the area of development, Dr. Mohr’s primary expertise is in designing applications that can be deployed to phones and desktop computers aimed at treating mental health disorders. While many of these have been relatively standard applications, he is also developing methods of harnessing sensor data from the phone to identify user states that are relevant to the treatment of depression. A second area of development focuses on developing applications aimed at improving adherence to medications and medical regimens. These applications are being deployed in General Internal Medicine, Community Health Centers, and Psychiatry. Finally, Dr. Mohr examines methods of implementing behavioral intervention technologies in the healthcare settings. In general, behavioral intervention technologies are not effective in improving symptoms when delivered as standalone treatments. Dr. Mohr has developed and evaluated methods of providing low intensity coaching support to enhance the use and effectiveness of behavioral intervention technologies. These coaching models can use health professionals, lay people, and peers.
He is also interested in the relationship between stress, depression and inflammation, particularly in multiple sclerosis.
For more information visit the faculty profile of David Mohr, PhD.
Publications
See Dr. Mohr's publications in PubMed.
Contact

Luisa Morales-Nebreda
Assistant Professor, Medicine (Pulmonary and Critical Care)
Immune mechanisms in lung injury and repair

Luisa Morales-Nebreda
Research Area(s): Computational Biology and Bioinformatics; Immunology; Lung Biology
My research program focuses on determinants of recovery from viral pneumonia in aged hosts. In the context of viral-induced lung injury, my laboratory aims to identify novel inter-cellular signaling pathways between immune cell subsets and the injured alveolar endothelium that promote tissue repair throughout the lifespan. Our group uncovered that during recovery from influenza pneumonia, age determines the tissue protective function of regulatory T (Treg) cells in a cell-autonomous manner (JCI Insight, 2021). Moreover, we recently identified distinctive T cell responses as correlates of immune protection and predictors of clinical outcomes in patients with respiratory failure from severe SARS-CoV-2 pneumonia (Nat Immunol, 2024). Stemming from these observations, we are interested in elucidating how T cells orchestrate endothelial cell repair and accelerate resolution of lung inflammation in aged hosts. To address these questions, we leverage multi-parameter flow cytometry, microscopy-based imaging and multi-omic technologies (sc-RNA-seq and spatially-resolved transcriptomics) to investigate the molecular-level underpinnings of cell-cell interaction biology in our innovative murine models and human samples.
For lab information and more, see Dr. Morales-Nebreda's faculty profile.
Publications
See Dr. Morales-Nebreda's publications on PubMed.
Contact
Contact Dr. Morales-Nebreda

William A Muller
Professor, Pathology (Experimental Pathology), Pathology (Autopsy)
Emigration of leukocytes across vascular endothelial cells in the process of inflammation

William A Muller
Research Area(s): Cancer Biology; Cardiovascular and Renal Biology; Cell Biology; Immunology
Most diseases are due to or involve a significant component of inflammation. My lab studies the inflammatory response at the cellular and molecular level. We are focused on the process of diapedesis, the "point of no return" in inflammation where leukocytes squeeze between tightly apposed endothelial cells to enter the site of inflammation. We have identified and cloned several molecules that are critical to the process of diapedesis (PECAM (CD31), CD99, and VE-cadherin) and are studying how they regulate the inflammatory response using in vitro and in vivo models. We have recently described the Lateral Border Recycling Compartment, a novel para-junctional organelle that contains PECAM and CD99 and is critical for diapedesis to occur. We are currently investigating how this compartment regulates diapedesis in the hope of finding novel and highly specific targets for anti-inflammatory therapy.
The “holy grail” of therapy is to develop selective anti-inflammatory agents that block pathologic inflammation without interfering with the body’s ability to fight off infections or heal wounds. By understanding how endothelial cells at the site of inflammation regulate leukocyte diapedesis, we are hoping to do just that. We have identified several molecules critical for diapedesis in acute and chronic inflammatory settings that can be genetically deleted or actively blocked to markedly inhibit clinical symptoms (e.g. in a mouse model of multiple sclerosis) and tissue damage (e.g. in a mouse model of myocardial infarction) without impairing the normal growth, development, and health of these mice. Our inflammatory models include atherosclerosis, myocardial infarction, ischemia/reperfusion injury, stroke, dermatitis, multiple sclerosis, peritonitis, and rheumatoid arthritis. We are also using 4-dimensional intravital microscopy to view the inflammatory response in real time in living animals.
Basic questions/issues that the work seeks to address:
- What are the molecular mechanisms and signaling pathways that endothelial cells use to regulate the inflammatory response?
- How can we therapeutically treat inflammatory diseases without compromising the ability of the immune system to respond to new threats?
- Do circulating tumor cells use the same mechanisms as leukocytes to cross blood vessels when they metastasize?
For more information, visit the faculty profile of William A Muller, MD, PhD
Publications
View Dr. Muller's publications at PubMed
Contact

Hidayatullah G Munshi
Professor, Medicine (Hematology and Oncology)
Role of fibro-inflammatory stromal reaction in pancreatic cancer progression

Hidayatullah G Munshi
Research Area(s): Cancer Biology; Pharmacology
The Munshi Lab is focused on understanding the role of the key collagenase MT1-MMP (MMP-14) and members of the Snail family transcription factors in pancreatic cancer progression using transgenic mouse models. We are also interested in understanding how the pronounced fibrotic reaction induces epigenetic changes to contribute to chemotherapy resistance. We have shown that the collagen microenvironment induces histone acetylation and that targeting 'readers' of histone acetylation marks using BET inhibitors can limit growth of pancreatic cancer cells. We plan to evaluate the efficacy of BET inhibitors in our mouse models with the eventual goal of testing this class of inhibitors in patients with pancreatic cancer.
For more information, visit the faculty profile page of Hidayatullah G Munshi, MD.
Publications
View lab publications via PubMed.
Contact Us
Contact Dr. Munshi at 312-695-6180 or the Munshi Lab at 312-503-0312.

Satish N Nadig
Professor, Surgery (Organ Transplantation), Microbiology-Immunology, Pediatrics
Novel Organ Preconditioning Strategies in Solid Organ Transplantation

Satish N Nadig
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Immunology; Metabolism and Endocrinology; Pharmacology
Dr. Nadig directs the NIH-funded Comprehensive Transplant Immunobiology Laboratory (CTIL) which is dedicated to investigating novel innovations in solid organ transplantation. Our research focus includes all aspects of transplantation spanning donor organ management to allograft tolerance induction. In addition, we are also investigating techniques that will lead to the next era of transplantation, including bioengineering and targeting cell-specific immunometabolism. The overarching aim of the laboratory is to pre-condition organs prior to implantation to mitigate or eliminate the necessity for patients to take systemic immunosuppressive medications. These pre-conditioning strategies involve nanotherapeutic-based targeted immunoppressant delivery to organs themselves, bolstering endothelial cell health in the preservation phase of organ transplantation, and manipulating the immunometabolism of organs ex vivo to render them less immunogenic upon implantation, to name a few.
For more information, see Dr. Nadig's faculty profile.
Publications
See Dr. Nadig's publications on PubMed.
Contact

Mojgan H Naghavi
Professor, Microbiology-Immunology
Microtubule regulation and function during infection by Human Immunodeficiency Virus (HIV)

Mojgan H Naghavi
Research Area(s): Cell Biology; Microbiology; Neurobiology
Our research focuses on infection by Human Immunodeficiency Virus type 1 (HIV-1), a retrovirus and causative agent of acquired immunodeficiency syndrome (AIDS). In addition to suppressing the immune system, rendering victims susceptible to opportunistic infections, HIV-1 can cross the blood-brain barrier and cause serious damage to the central nervous system, ultimately leading to HIV-associated dementia.
We are interested in how HIV-1 particles move within infected cells, including brain cell types such as microglia. Our work focuses on how the virus exploits host microtubules, the intracellular filaments that mediate cargo trafficking to different subcellular sites within the cell.
Our earlier work, employing a variety of screening approaches, identified a number of host proteins involved in cytoskeletal regulation and motor function as playing key roles in the early stages of HIV-1 infection. This includes Ezrin-Radixin-Moesin (ERM) proteins, which cross-link the actin and microtubule cytoskeletons. In exploring their role in HIV-1 infection, we identified the first biological function for the host protein, PDZD8, demonstrating that it binds ERMs to control microtubule stability. Furthermore, we uncovered that PDZD8 is a direct target for the HIV-1 protein, Gag.
Other work in our laboratory has shown that HIV-1 can induce the formation of highly stable microtubule subsets to facilitate early HIV-1 trafficking to the nucleus. We are interested in the role played in this process by proteins such as PDZD8, as well as a family of specialized microtubule regulatory proteins called +TIPs, which accumulate at the ends of dynamically growing microtubule filaments to control their growth and stability. We are also interested in the function of microtubule motors and cargo adaptor proteins in HIV-1 infection. In particular, we are exploring how Fasciculation and Elongation factor Zeta-1 (FEZ-1), a kinesin-1 adaptor protein that is highly expressed in neurons, functions to control HIV-1 infection. Our work employs a range of approaches, including biochemical characterization of protein-protein interactions as well as live imaging of fluorescently-labeled HIV-1 particles as they move within infected cells.
The ultimate goal of our work is to understand the molecular basis behind how microtubules, regulators of microtubule dynamics and microtubule motor proteins function to enable HIV-1 movement to and from the nucleus.
For lab information and more, see Dr. Naghavi's faculty profile and lab website.
Publications
See Dr. Naghavi's publications at PubMed.
Contact
Contact Dr. Naghavi at 312-503-4294.

Andrew M Naidech
Professor, Neurology (Neurocritical Care), Medical Social Sciences (Outcome and Measurement Science), Preventive Medicine (Biostatistics and Informatics), Anesthesiology, Neurological Surgery, Neurology (Stroke and Vascular Neurology)
Devise new treatments, improve protocols and maximize quality of life for patients with acute neurological injuries

Andrew M Naidech
Research Area(s): Computational Biology and Bioinformatics; Neurobiology
The section of neurocritical care combines researchers working on complementary areas of neurological emergencies. Ongoing projects include cerebrovascular autoregulation and cognition (Farzaneh Sorond, MD, PhD), patient-centered outcomes research, translational research for acute intracerebral hemorrhage such as acute correction of platelet activity (Andrew Naidech, MD, MSPH), sleep dysfunction and circadian rhythms in acute illness (Matthew Maas, MD), cerebral edema and hepatic encephalopathy (Eric Liotta, MD), and basic neuroscience (Minjee Kim, MD).
Publications
See Dr. Naidech's publications on PubMed.
Contact
Andrew Naidech, MD, MSPH, FANA
Professor of Neurology

Diana Nora Vaz Saleiro
Assistant Professor, Medicine (Hematology and Oncology)
Uncovering and targeting novel resistance mechanisms to cancer immunotherapies

Diana Nora Vaz Saleiro
Research Area(s): Cancer Biology; Immunology
The Saleiro Lab is interested in basic and translational research on regulation of resistance mechanisms to immunotherapies, focusing on melanoma and myeloproliferative neoplasms (MPNs). Our goal is to translate these findings into personalized targeted and immunotherapies, contributing to increasing cancer patients’ response rates and survival. Our projects focus on identifying and characterizing novel targetable molecules within the interferon (IFN) pathways to enhance IFN-mediated anti-cancer effects. IFNs are considered potent cytokines with antiviral, antiproliferative, immunomodulatory, and antineoplastic properties and key regulators of responses to immunotherapies. However, the molecular mechanisms of IFN action remain largely elusive, complicating the overall understanding of their biological effects and precluding advances on their clinical use/regulation. In past studies, we demonstrated that the kinase ULK1 plays a unique role in controlling both type I (IFNa/b) and type II IFN (IFNg) responses, independent of its role in autophagy. Through proteomic and molecular biology analyses, we have identified novel clinically targetable ULK1 protein complexes for the treatment of melanoma and MPNs. These results define previously unknown and distinct IFN pathways that appear to be key elements required to control IFN-mediated immune and antineoplastic responses. Current studies are aimed at increasing our understanding of interferons' actions and of the kinase ULK1 to support the development of new and more efficient targeted-therapies and immunotherapies involving the modulation of IFN-mediated responses. The disease models used in the lab are melanoma and myeloproliferative neoplasms. Specific research areas include regulation of interferon signaling in tumor and immune cells, identifying and characterizing novel immunosuppressive proteins, evaluating and characterizing novel therapeutic strategies in preclinical cancer mouse models, and performing immune cell-tumor cell killing studies using state-of-the art technologies.
For more information view Dr. Saleiro's faculty profile.
Publications
Please see Dr. Saleiro's publications on PubMed.
Contact Information
Contact Dr. Saleiro

Guillermo Oliver
Professor, Medicine (Nephrology and Hypertension)
Acquisition of specific and unique morphological and functional characteristics of cells and organs during embryogenesis

Guillermo Oliver
Research Area(s): Developmental and Stem Cell Biology; Neurobiology
The Oliver Lab focuses on understanding how each specific cell type and organ acquires all its specific and unique morphological and functional characteristics during embryogenesis. Alterations in the cellular and molecular mechanisms controlling organ formation can result in major defects and pathological alterations. Our rationale is that a better knowledge of the basic processes controlling normal organogenesis will facilitate our understanding of disease. Our goal is to dissect the specific stepwise molecular processes that make each organ unique and perfect. Our major research interests are the forebrain, visual system and the lymphatic vasculature and to address those questions we use a combination of animal models and 3D organ culture systems, stem cells and iPS cells.
Related to the lymphatic vasculature, our lab identified years ago the first specific marker for lymphatic endothelial cells and generated the first mouse model devoid of lymphatics. We have characterized many of the critical steps leading to the formation of the lymphatic vasculature. We have also reported that a defective lymphatic vasculature can cause obesity in mice and we are currently trying to determine whether this is also valid in humans.
In case of the central nervous system, our focus is to characterize how complex structures such as the forebrain and eye are formed. For that we have started to apply 3D organ culture systems derived from stem cell and iPS that allow us to grow eyes in a petri dish. Using this approach we expect to dissect the genes and mechanisms controlling these developmental processes.
For more information, visit Dr. Oliver’s faculty profile or visit the Guillermo Oliver Lab Site.
Publications
View Dr. Oliver's publications at PubMed
Contact
Phone 312-503-1651
Puneet Opal
Professor, Neurology (Movement Disorders), Cell and Developmental Biology
Cellular basis of neurodegeneration
Puneet Opal
Research Area(s): Biochemistry and Structural Biology; Genetics, Epigenetics and Genomics; Neurobiology
The long-term goal of my laboratory is to understand the cellular basis of neurodegeneration. We are testing the idea that neurodegeneration results from derangements in relatively few but strategic sub-cellular pathways. By identifying critical components of these pathways one could begin to not only understand the biology of neurodegeneration, but also embark on the design of novel therapeutic agents.
We are currently studying the autosomal dominant disorder Spinocerebellar Ataxia Type 1 (SCA1), a relentless disease that affects cerebellar Purkinje cells and brainstem neurons. This disorder is caused by a polyglutamine expansion in the involved disease protein and is thus similar to a growing number of disorders, including Huntington disease, that share this mutational mechanism. Patients with SCA1 begin to display cerebellar signs characterized by motor incoordination or ataxia in early to mid adulthood. Unfortunately there is no treatment for this disease and patients eventually succumb from the complications of brainstem dysfunction.
Current Projects
-
Testing the transcriptional hypothesis in SCA1 pathogenesis:
One of the earliest features of this disease is change in the gene expression signature within neurons affected in this disease. We are elucidating the pattern of gene expression changes in the vulnerable Purkinje cell population and identifying the contribution of these alterations to pathology.
-
Testing the role of the vascular and angiogenic factor VEGF in SCA1 pathogenesis.
One of the genes that we have already found to be down-regulated is the neurotrophic and angiogenic factor VEGF. Importantly, we have discovered that genetic or pharmacologic replenishment of VEGF mitigates SCA1 pathogenesis. These results suggest a novel therapeutic strategy for this incurable disease and a possible cross-talk between the degenerating cerebellum and its microvasculature. We are pursuing mechanistic experiments to learn how low VEGF mediates SCA1 pathology. In addition, we are working actively towards testing the potential for VEGF as a therapy in human ataxic disorders.
-
Testing the role of ataxin-1 misfolding and clearance in disease pathogenesis.
Several studies suggest that ataxin-1 accumulates in neurons because of its inability to be cleared by the protein-misfolding chaperone pathway. We are testing different strategies to promote ataxin-1 clearance.
In addition to spinocerebellar ataxia, we are also studying genetic parkinsonian and dystonic syndromes.
For more information see the faculty profile of Puneet Opal, MD, PhD or visit the Opal Lab website.
Publications
View Dr. Opal's full list of publications at PubMed.
Contact
Phone: 312-503-4699

Ertugrul Ozbudak
Professor, Cell and Developmental Biology
Biological form of “art”: Pattern formation in developing tissues and organs

Ertugrul Ozbudak
Research Area(s): Cell Biology; Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics
Our lab’s overriding interest is to achieve a mechanistic understanding of embryonic development and pattern formation by integrating quantitative experiments with computational modeling. In our lab, we combine single-cell confocal microscopy measurements are combined with time-resolved perturbation experiments, genome-wide techniques, biophysical modeling and computational simulations. Our aspiration is to discover fundamental principles of development, discern the etiology of birth defects and inform strategies for regenerative medicine. We are currently investigating this goal in two different systems: somite segmentation and fin/limb development. We will soon be expanding our work to other tissues and organs by using both zebrafish embryos and human organoids.
For more information see the Dr. Ozbudak's faculty profile.
Publications
View Dr. Ozbudak's full list of publications at PubMed.
Contact

Hande Ozdinler
Associate Professor, Neurology (Neuromuscular Disease)
Cortical component of motor neuron circuitry degeneration in amyotrophic lateral sclerosis (ALS) and other related disorders

Hande Ozdinler
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Neurobiology; Pharmacology
We are interested in the cellular and molecular mechanisms that are responsible for selective neuronal vulnerability and degeneration in motor neuron diseases. Our laboratory especially focuses on the corticospinal motor neurons (CSMN) which are unique in their ability to collect, integrate, translate and transmit cerebral cortex's input toward spinal cord targets. Their degeneration leads to numerous motor neuron diseases, including amyotrophic lateral sclerosis, hereditary spastic paraplegia and primary lateral sclerosis.
Investigation of CSMN require their visualization and cellular analysis. We therefore, generated reporter lines in which upper motor neurons are intrinsically labeled with eGFP expression. We also characterized progressive CSMN degeneration in various mouse models of motor neuron diseases and continue to generate reporter lines of disease models, in which the upper motor neurons express eGFP.
The overall goal in our investigation, is to develop effective treatment strategies for ALS and other related motor neuron diseases. We appreciate the complexity of the disease and try to focus the problem from three different angles. In one set of studies, we try to reveal the intrinsic factors that could contribute to CSMN vulnerability by investigating the expression profile of more than 40,000 genes and their splice variations at different stages of the disease. In another set of studies, we try to understand the role of non-neuronal cells on motor neuron vulnerability and degeneration, using a triple transgenic mouse model, in which the cells that initiate innate immunity are genetically labeled with fluorescence in an ALS mouse model. These studies will not only reveal the genes that show alternative splice variations, but also inform us on the canonical pathway and networks that are altered with respect to disease initiation and progression.
Even though the above mentioned studies, which use pure populations of neurons and cells isolated by FACS mediated approaches, will reveal the potential mechanisms that are important for CSMN vulnerability, it is important to develop therapeutic interventions. One of the approach we develop is the AAV-mediated gene delivery directly into the CSMN via retrograde transduction. Currently, we are trying to improve CSMN transduction upon direct cortex injection.
Identification of compounds that support CSMN survival is an important component of pre-clinical testing. We develop both in vitro and in vivo compound screening and verification platforms that inform us on the efficiency of compounds for the improvement of CSMN survival.
In summary, we generate new tools and reagents to study the biology of CSMN and to investigate both the intrinsic and extrinsic factors that contribute to their vulnerability and progressive degeneration. We develop compound screening and verification platforms to test their potency on CSMN and develop AAV-mediated gene delivery approaches. Our research will help understand the cellular basis of CSMN degeneration and will help develop novel therapeutic approaches.
For more information see the faculty profile of Pembe Hande Ozdinler, PhD or the Ozdinler Lab website.
Visit the Les Turner ALS Center
Publications
View Dr. Ozdinler's full list of publications at PubMed.
Contact
Phone: 312-503-2774

Joo-Seop Park
Professor, Medicine (Nephrology and Hypertension)
Kidney development and disease: How cell identities are determined by signaling pathways and transcription factors

Joo-Seop Park
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics
The Park lab studies how stem/progenitor cells differentiate into specific cell types using the mouse kidney as a model system. The nephron, the functional unit of the kidney, is composed of at least 15 distinct cell types. Since all of the cell types found in the nephron originate from the common nephron progenitor cells, the mouse kidney serves as an excellent system to study cell fate decisions of stem/progenitor cells. We aim to (1) determine the roles of developmental signaling pathways in nephron formation, (2) identify transcription factors that define cell identities for each cell type found in the nephron, and (3) elucidate how these transcription factors coordinate with signaling pathways in gene regulatory networks.
For lab information and more, see Dr. Park's faculty profile.
Publications
See Dr. Park's publications on PubMed.
Contact
Contact Dr. Park

Jones G Parker
Assistant Professor, Neuroscience, Pharmacology, Psychiatry and Behavioral Sciences
Targeting neural substrates to improve treatment outcomes for neuropsychiatric diseases

Jones G Parker
Research Area(s): Neurobiology; Pharmacology
We use imaging approaches to acquire large-scale recordings of neural activity during behavior, focusing on deep-brain areas implicated in neurological and psychiatric diseases, such as striatum. We use these tools to delineate the functional contribution of neuronal sub-populations in these brain areas to normal behavior in control subjects and pathological behavior in models for brain diseases. Our aim is to target these neural substrates to improve treatment outcomes for neuropsychiatric diseases.
For lab information and more, see Dr. Parker's faculty profile and lab website.
Publications
See Dr. Parker's publications on PubMed.
Contact
Contact Dr. Parker at 312-503-3165.

Saba Parvez
Assistant Professor, Cell and Developmental Biology
Interrogating how the poorly characterized genome regulates animal development

Saba Parvez
Research Area(s): Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics
The functional roles of a vast majority of the genome remains poorly characterized. Large-scale genetic screens, primarily in tissue cultures, have been instrumental in uncovering the functions of the poorly characterized regions. However, in vitro cell cultures are phenotypically limited to observations at the level of single cells, monolayers, or organoids. Our lab applies high throughput approaches in zebrafish to systematically perturb various poorly characterized genomic regions to understand their roles in regulating animal development and behavior. In particular, we are keen on understanding how certain highly conserved noncoding genomic regions regulate developmental processes. These explorations are complemented by the development of novel technologies to perturb the genome and new ways of measuring phenotypic changes resulting from the perturbation. Students joining the lab will have the opportunity to train in bioinformatics and genomic technologies, CRISPR-based genetic perturbation methods, high content morphological and behavioral phenotyping approaches, cellular and molecular biology techniques, and state-of the art imaging platforms.
For lab information and more, see Dr. Parvez's faculty profile.
Publications
See Dr. Parvez's publications on PubMed.
Contact
Contact Dr. Parvez

Clara B Peek
Assistant Professor, Biochemistry and Molecular Genetics, Medicine (Endocrinology, Metabolism and Molecular Medicine)
Circadian clock control of fuel selection and response to nutrient stress

Clara B Peek
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology
The Peek Lab is focused on understanding the interplay between hypoxic and circadian transcriptional pathways both at the genomic and nutrient signaling levels. Peek aims to uncover novel mechanisms linking circadian clocks to the control of metabolic function and disease, such as type 2 diabetes and cancer. The lab utilizes metabolic flux analyses, in vivo metabolic and behavior monitoring, and next-generation sequencing in their research.
For lab information and more, see Dr. Peek's faculty profile and lab website.
Publications
See Dr. Peek's publications on PubMed.
Contact
Contact Dr. Peek at 312-503-6973.

Minoli Perera
Associate Professor, Pharmacology
Pharmacogenomics research in minority patient populations

Minoli Perera
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Pharmacology

The Perera laboratory focuses on pharmacogenomics (using a patient's genome to predict drug response) in minority populations. Working in this translation research space requires both clinical expertise as well as the use of high-throughput basic science approaches. Our goal is to bring the benefits of precision medicine to all US populations.
The Perera lab has recruited patient populations from around the world. The data collection includes genomic (DNA), transcriptomic (mRNA), pharmacokinetic and clinical data. We then integrate these different data sources to understand genetic drivers of drug response (e.g. genetic predictors of adverse events) as well as disease. By studying minority populations the lab has discovered genetic risk variants that may benefit the implementation of precision medicine in African Americans and others.
Recent Findings
- Warfarin Bleeding Risk Association study
We recently discovered a genetic variant that predispose African Americans to bleeding complications while on anticoagulant drugs. These bleeds occurred even when the patient was within the therapeutic window for the medication. We hope that this new data will help to identify high risk individuals prior to therapy. - Novel African-specific genetic polymorphisms predict the risk of venous thromboembolism
We discovered a new genetic variant associated with a 2.5 fold increase in risk of developing a blood clot. We went on to show that this SNP significantly affects the expression of a key protein in the coagulation cascade. View article on PubMed. - Common genetic variant is predictive of warfarin metabolism and gene expression in African Americans
We tested the association of a SNP, previously shown to effect gene expression CYP2C9, for association with warfarin drug clearance (pharmacokinetics). This SNP increased the expression of CYP2C9 (enzyme that metabolized warfarin), hence causing fast clearance of the drug. This African American-specific SNP may help to explain the higher warfarin dose required by African Americans in general. View article on PubMed.
Current Projects
- Genomics of Drug Metabolism
We are using African America primary hepatocytes to understand the genetic regulation of drug metabolizing enzymes that are involved in a majority of drug used in the US. - Anticoagulant Pharmacogenomics
We are conducting several genetic association studies to understand both the genetic drivers and the biological mechanisms behind response and adverse effect to anticoagulant medications. - Pharmacogenomics of Inflammatory Bowel disease
We are investigating the genetic predictors of primary non-response to biologic therapies used in inflammatory bowel disease. Studies have implication for other autoimmune disorders that target the same pathways. - eMERGE
We are involved in analyzing the GWAS and sequencing data specifically for genomics variation affect key pharmacogenomics gene in African Americans.
For lab information and more, see Dr. Perera's faculty profile and lab website.
Publications
See Dr. Perera's publications on PubMed.
Contact
Contact Dr. Perera at 312-503-6188 or the lab at 312-503-4119.

Harris R Perlman
Professor, Medicine (Rheumatology)
Impact of macrophages in pathogenesis of rheumatic disease

Harris R Perlman
Research Area(s): Immunology
Macrophages have emerged as key players in the development of inflammation and fibrosis in central target organs including the synovium, kidney and lung during the pathogenesis and remission of rheumatoid arthritis (RA), systemic lupus erythematosus (SLE) and systemic sclerosis (SSc), respectively. Macrophages also contribute to the co-morbidities associated with these diseases including atherosclerosis and obesity. We observed marked heterogeneity in the macrophage population within diseased tissues that is dependent on their origin (embryonic vs. bone marrow derived), target organ and microenvironment. Moreover, these macrophages are extremely plastic and can alter their phenotype throughout the course of disease. Based on our data we developed a central hypothesis that during the initiation and early progression phase of disease tissue resident macrophages that normally function to maintain tolerance to local antigens, are overwhelmed by recruited pro-inflammatory or pro-fibrotic monocyte derived macrophages or pro-inflammatory dendritic cells depending on the target organ and environmental milieu. As the disease progresses to the chronic phase, however, the recruited macrophages acquire characteristics reminiscent of tissue resident macrophages while retaining a pro-inflammatory and pro-fibrotic phenotype, resulting in failed resolution of inflammation and progressive tissue destruction and fibrosis. The data anticipated from our projects would be the first to demonstrate a direct linkage of macrophage ontogeny and activation to disease activity and tissue damage. In addition, our studies allow us to explore commonalities in macrophage function between diseases that could lead to broad therapeutic interventions. In our state-of-the-art murine models we use cutting-edge technologies that we developed including micro-MRI, CT and SPECT to evaluate joint inflammation, bone destruction and lung fibrosis, Luminex-based gene arrays and multiparameter flow cytometry/sorting, whole population RNA seq and single cell RNA Seq and Chip-seq. We will cross-reference these data with those we will obtain through the AMP programs, which examine macrophage heterogeneity in the synovium and kidney from patients with rheumatic disease. This will allow us to rapidly move to functional analyses of relevant pathways and testing of new therapeutic strategies in the mouse models. I believe that our data has the potential to be paradigm shifting and transformative for the field of rheumatic disease.
For more information related to the Perlman Lab, or to connect with us, please see Harris R Perlman’s, PhD, profile.
Publications
View Dr. Perlman's publications at PubMed
Contact
Contact Dr. Perlman at 312-503-8003 or the Perlman Lab at 312-503-1933.

Marcus Ernst Peter
Professor, Medicine (Hematology and Oncology), Biochemistry and Molecular Genetics
Cell death including apoptosis, a fundamental process to regulate homeostasis of all tissues and to eliminate unwanted cells specifically in the immune system

Marcus Ernst Peter
Research Area(s): Cancer Biology; Cell Biology; Immunology
Another interest lies in the study of RNA interference and based on toxic RNAs to development a novel form of cancer treatment.
Publications
View lab publications via PubMed.
For more information, visit the faculty profile page of Marcus Peter, PhD or the laboratory's website.
Contact Us
Contact Dr. Peter at 312-503-1291 or the Peter Lab at 312-503-2883.

Leonidas C Platanias
Professor, Medicine (Hematology and Oncology), Biochemistry and Molecular Genetics
Signaling pathways in different types of cancers and development of novel therapies to specifically kill cancer cells

Leonidas C Platanias
Research Area(s): Cancer Biology; Cell Biology; Pharmacology
Cell signaling is part of an intricate system of events activated by various stimuli that coordinate cell responses. Our laboratory is interested in unveiling pathways involved in cancer development in order to target them and control cancer progression. For over two decades, Dr. Platanias’ laboratory has identified several cellular cascades activated by IFN, ATRA and arsenic. Our research on Type I IFN found an essential role for SKAR protein in the regulation of mRNA translation of IFN-sensitive genes and induction of IFN-α biological responses. We also provided evidence for unique function of mTORC2 complex in inducing Type I IFN response. Our studies on arsenic signaling revealed a direct binding of this compound to a kinase called AMPK as a mechanism underlying its anti-leukemic activity. Other work included the activation of biological responses by BCR-ABL oncoprotein through the mTOR pathway. Dr. Platanias’ laboratory is also involved in testing new compounds in combination with approved therapies in order to identify synergy and improve the risk/benefit ratios of current therapeutic regimens for patients.
For more information, visit the faculty profile page of Leonidas Platanias, MD, PhD.
Publications
View lab publications via PubMed.
Contact Us
Contact Dr. Platanias at 312-908-5250 or the Platanias Lab at 312-503-4500.

Seth M Pollack
Professor, Medicine (Hematology and Oncology)
Immunotherapy in sarcoma

Seth M Pollack
Research Area(s): Cancer Biology; Immunology
Sarcomas are rare cancers of the bone and connective tissues that are quite deadly and frequently impact adolescents and young adults. The Pollack Lab is a cancer immunotherapy lab focused on curing sarcoma through the development of better immunotherapy treatments. We focusing on manipulating the sarcoma tumor immune microenvironment to make better vaccines, T cell therapies and check point inhibitor strategies. These are the current, most-active main focuses of the lab.
1) Exploiting the naturally occurring canine model to improve gene-engineered cellular therapy: Creating accurate models of cancer is hard. Most labs use mouse models of cancer — and we do this too for many applications. However, it is very important to be cognizant of what these models can tell us about human disease and where their limits are. Our lab studies cancer immunotherapy and many mouse models of human cancers have immunodeficiencies. Other mouse models have important differences with respect to the tumor biology. Because our lab is always coming up with new ideas for improving sarcoma immunotherapy with a focus on bringing treatments to the clinic, we need a great model for their development.
While sarcomas are (thankfully) relatively rare in humans, these cancers are one of the most common cancers in pet dogs. There is very good science showing that these canine tumors very closely resemble their human counterparts, probably in part because they develop in a natural way and may have a tumor evolution that also mirrors human disease. These are normal dogs with healthy immune systems. We have developed many processes and techniques for studying canine cancers and immunotherapies using gene engineered therapies and this work is currently moving at full speed. We have also developed relationships with academic veterinarians, key non-profit organizations (https://www.ccralliance.org) and currently have two active, enrolling trials in the veterinary clinic testing new gene engineered immunotherapies in dogs. While our main goal is to answer key questions regarding therapy for human sarcomas, we are also trying to help the pets on these trials as well as their families.
2) Improving Cell Therapy for Synovial Sarcoma and Myxoid/Round Cell Liposarcoma: The early years of the Pollack Lab were entirely dedicated to tediously isolating, sorting, and expanding rare cancer specific T cells from the blood of cancer patients for analysis and therapy. As part of this work, we generated clinical grade protocols and ultimately cell-therapy sarcoma clinical trials with clinical responses that were unfortunately short-lived. We found that despite progression, these patients retained antigen expression and continued to have rare antigen presenting cells circulating in their peripheral blood. When we co-cultured isolated the rare cancer specific T cells from the patient’s blood with tumor lines in the lab, these cells would not kill tumor untreated however, after treatment with IL-15 they vigorously proliferated and efficiently killed targets. This is one of the phenomenon that we are very excited about exploring.
In correlative studies, we identified MHC down-regulation as an important means of immune evasion in these sarcoma. We established an investigator-sponsored trial supported, testing whether interferon gamma could up-regulate MHC in these tumors. That trial demonstrated the potential for interferon gamma to up-regulate MHC and recruit tumor specific T cells into the sarcoma immune microenvironment but also increased expression of PD-L1 in the tumor which led to a Cancer Immunotherapy Trials Network study of interferon gamma and anti-PD1 trial. This trial is complete and we are actively studying samples from this study; we believe this analysis will lead to fundamental insights that will alter future trials of both engineered T cells and vaccines in these terrible diseases.
3) Understanding the Changing T Cell Receptor Repertoire in the Sarcoma Immune Microenvironment: The human body has an incredible diversity of T cells capable of recognizing all sorts of threats, including infections and cancers. Our early work analyzing the T cell repertoire in sarcomas demonstrated that certain sarcomas have a more robust and focused T cell response. More recently, we have focused on understanding the changing T cell repertoire for sarcoma patients on clinical trials. We have identified key T cell receptor sequences in certain clinical situations that we believe have important clinical significance. We have also been leaders in characterizing the sarcoma immune microenvironment both in its natural state and as it changes on immunotherapy trials. Our lab is actively working to isolate T cells with these sequences and to better understand their phenotype and function.
For lab information and more, see Dr. Pollack's faculty profile.
Publications
See Dr. Pollack's publications on PubMed.
Contact
Contact Dr. Pollack.

Richard M Pope
Professor Emeritus, Medicine (Rheumatology)
Biology of macrophages in the pathogenesis of rheumatoid arthritis (RA)

Richard M Pope
Research Area(s): Cell Biology; Immunology
Our laboratory has identified the upregulation of the anti-apoptotic protein, Flice Like Inhibitory Protein (FLIP), during monocyte to macrophage differentiation. They have demonstrated that FLIP is highly expressed in the rheumatoid joint and is responsible for protecting RA macrophages from Fas-mediated apoptosis. These studies have been extended to examine the in vivo relevance of FLIP to macrophage biology. Mice with FLIP conditionally deleted in myeloid cells are not capable of developing macrophages. The relevance of these observations to chronic inflammatory arthritis is currently under investigation. Since macrophages are critical to the pathogenesis of RA, future studies will focus on macrophage specific FLIP as a therapeutic target in RA.
Additional studies in the laboratory are focusing on the role of endogenous TLR ligands as potential contributors to the persistent activation of macrophages in the RA joint. The Pope laboratory has identified an endoplasmic reticulum (ER) localized protein called gp96, which binds to TLRs within ER of macrophages and correctly transports them to the cell membrane or endosome. In patients with RA, gp96 is highly increased in RA synovial tissue, particularly in macrophages, and is found in RA synovial fluid in high concentrations. gp96 binds to the extracellular domains of TLR2 and TLR4. Both recombinant gp96 and gp96 present in the RA synovial fluid is capable of activating TLR2 and to a lesser degree TLR4. Ongoing studies in the laboratory are further characterizing the mechanisms by which gp96 and other endogenous TLR ligands might contribute to the pathogenesis of RA employing in vitro studies utilizing cells isolated from the joints of patients with RA and experimental murine models of RA
Additional studies are ongoing in the laboratory to further identify and define the potential clinical relevance of endogenous TLR ligands in the RA joint employing three approaches which are dependent upon binding to endogenous ligands to TLRs. These approaches include the use of recombinant IgG Fc-TLR2 and IgG Fc-TLR4 to pull down TLR ligands from cells isolated from the RA joint; the use of HEK-TLR2 and HEK-TLR4 cells to bind endogenous TLR ligands in RA synovial fluid which will be identified employing a proteomics approach and utilizing a yeast 2 hybrid system where mRNA from inflammatory RA synovial tissue has been employed to develop the bait, while the extracellular domains to TLR2 and TLR4 are being used as the prey. Each of these approaches have identified candidate molecules which are being further characterized for a potential role in the pathogenesis of RA.
For more information related to the Pope Lab’s work, please see Richard M. Pope’s, MD, profile.
Publications
View Dr. Pope's publications at PubMed
Contact
Contact Dr. Pope at 312-503-8003 or the Pope Lab at 312-908-1965.

Murali Prakriya
Professor, Pharmacology, Medicine (Allergy and Immunology)
Calcium signaling, inflammation, and brain function

Murali Prakriya
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Immunology; Pharmacology
Research in the laboratory of Murali Prakriya, PhD, is focused on the molecular and cellular mechanisms of intracellular calcium (Ca2+) signaling. Ca2+ is one of the most ubiquitous intracellular signaling messengers, mediating many essential functions including gene expression, chemotaxis and neurotransmitter release. Cellular Ca2+ signals generally arise from the opening of Ca2+-permeable ion channels, a diverse family of membrane proteins. We are studying Ca2+ signals arising from the opening of store-operated Ca2+ channels (SOCs). SOCs are found in the plasma membranes of virtually all mammalian cells and are activated through a decrease in the calcium concentration ([Ca2+]) in the endoplasmic reticulum (ER), a vast membranous network within the cell that serves as a reservoir for stored calcium. SOC activity is stimulated by a variety of signals such as hormones, neurotransmitters and growth factors whose binding to receptors generates IP3 to cause ER Ca2+ store depletion.
The best-studied SOC is a sub-type known as the Ca2+ release activated Ca2+ (CRAC) channel encoded by the Orai1 protein. CRAC channels are widely expressed in immune cells and generate Ca2+ signals important for gene expression, proliferation and the secretion of inflammatory mediators. Loss of CRAC channel function due to mutations in CRAC channel genes leads to a devastating immunodeficiency syndrome in humans. Our goals are to understand the molecular mechanisms of CRAC channel activation, and their physiological roles especially in the microglia and astrocytes of the brain, and in the airway epithelial cells of the lung.
Recent Findings
Despite the fact that CRAC channels are found in practically all cells, their properties and functions outside the immune system remain largely unexplored. In order to fill this gap, we have begun investigation of CRAC channel properties and their functions in two major organ systems: in the brain and the lung.
- In the brain, we are studying the role of CRAC channels for dendritic Ca2+ signaling in excitatory neurons of the hippocampus, and their role in synaptic plasticity and cognitive functions. We have found that CRAC channels formed by Orai1 are critical for amplifying glutamate receptor evoked calcium signals in dendritic spines of hippocampal neurons, and this step is essential for driving structural and functional measures of synaptic plasticity and cognitive processes involving learning and memory.
- In a second project, we are studying the role of CRAC channels in driving neuroinflammation. We have found that CRAC channels formed by Orai1 are essential for the production and release of proinflammatory cytokines and chemokines in microglia and astrocytes. We are examining the relevance of this pathway for mediating inflammatory and neuropathic pain.
- A third project is examining the role of CRAC channels for mediating pro- and anti-inflammatory processes in the lung. We have found that CRAC channels are a major mechanism for mobilizing Ca2+ signals in lung epithelial cells, and the downstream production of both pro- and anti-inflammatory mediators. We are examining the relevance of this signaling for lung inflammation in the context of asthma.
For lab information and more, see Dr. Prakriya’s faculty profile and lab website.
Publications
See Dr. Prakriya's publications on PubMed.
Contact
Contact Dr. Prakriya at 312-503-7030.

Arthur B Prindle
Associate Professor, Biochemistry and Molecular Genetics, Microbiology-Immunology
Associate Professor, McCormick School of Engineering
Synthetic biology in microbial communities

Arthur B Prindle
Research Area(s): Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Microbiology; Pharmacology
The Prindle lab is interested in understanding how molecular and cellular interactions give rise to collective behaviors in microbial communities. While bacteria are single celled organisms, we now understand that most bacteria on our planet reside in the context of structured multicellular communities known as biofilms. However, most bacterial research is still performed on domesticated lab strains in well-mixed conditions. We simply do not know enough about the biology and behavior of the most pervasive life form on our planet. It is our goal to discover and understand these behaviors so that we may apply our understanding to engineer biomolecular systems as solutions to challenging biomedical problems, such as antibiotic resistance. To do this, we also work on developing technologies that can characterize collective metabolic and electrochemical dynamics that emerge in the context of biofilms.
For more information, see Dr. Prindle's lab website.
Publications
See Dr. Prindle’s publications on PubMed.
Contact
Contact Dr. Prindle

Susan E Quaggin
Professor, Medicine (Nephrology and Hypertension)
Molecular mechanisms of diabetic vascular complications, thrombotic microangiopathy, glomerular diseases and glaucoma

Susan E Quaggin
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Developmental and Stem Cell Biology; Pharmacology
Our lab focuses on the basic biology of vascular tyrosine kinase signaling in development and diseases of the blood and lymphatic vasculature. Our projects include uncovering the molecular mechanisms of diabetic vascular complications, thrombotic microangiopathy, glomerular diseases and glaucoma. Utilizing a combination of mouse genetic, cell biologic and proteomic approaches, we have identified key roles for Angiopoietin-Tie2 and VEGF signaling in these diseases. Members of the lab are developing novel therapeutic agents that target these pathways.
For more information, please see the faculty profile of Susan Quaggin, MD
Publications
See Dr. Quaggin's publication in PubMed
Contact

Karen M Ridge
Professor, Medicine (Pulmonary and Critical Care), Cell and Developmental Biology
Role of intermediate filaments in lung pathophysiology

Karen M Ridge
Research Area(s): Cancer Biology; Cell Biology; Lung Biology
The cytoskeletal protein vimentin plays a key role as a scaffold for the formation and activation of intracellular protein complexes. One such complex is the NLRP3 inflammasome, which assembles in response to danger signals such as influenza A virus or ATP released by damaged cells to produce mature IL-1β and IL-18. These inflammatory cytokines can induce lung injury, which can lead to fibrosis. One of our current goals is to pinpoint vimentin’s role in inflammasome assembly and activation.
Vimentin is also involved in all stages of cancer development, from PI3K/AKT and Erk pathway regulation in tumorigenesis, to its defining role in epithelial-to-mesenchymal transition, to metastatic cell invasion and migration, making it an intriguing therapeutic target. Our purpose in examining vimentin’s role in lung cancer is to determine whether its inhibition might be of benefit to patients.
To learn more, please visit the faculty profile pages of Karen M. Ridge, PhD or the Ridge Lab Website
Publications
View our lab’s publications in PubMed.
Contact Us
Email Dr. Ridge
Phone 312-503-1648 or the Ridge Lab at 312-503-0403

Gabriel J Rocklin
Assistant Professor, Pharmacology
High-throughput methods for protein biophysics and protein design, with a focus on protein therapeutics

Gabriel J Rocklin
Research Area(s): Biochemistry and Structural Biology; Pharmacology
Key questions include: How do protein sequence and structure determine folding stability, conformational dynamics, and resistance to aggregation/degradation-inducing stresses? Can we quantitatively predict these protein "phenotypes" from genotype (sequence) using computational modeling? How do we design protein therapeutics that optimize these phenotypes for a particular application? To answer these questions, we combine large-scale de novo computational protein design with high-throughput methods such as display selections, mass spectrometry proteomics, and next-generation sequencing, enabling us to test thousands of proteins in parallel. By combining these technologies, we seek to develop efficient "design-test-analyze" cycles, iterating our way to an improved, quantitative understanding of protein biophysics and more advanced protein therapeutics.
For lab information and more, see Dr. Rocklin's faculty profile and lab website.
Publications
See Dr. Rocklin's publications on PubMed.
Contact
Contact Dr. Rocklin at 312-503-4892.

Smriti Sangwan
Assistant Professor, Pharmacology
Molecular basis of protein quality control

Smriti Sangwan
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Pharmacology
Protein quality control is critical for cellular health and is constantly monitored via signaling systems to ensure that all proteins fold correctly and are delivered to their designated cellular location. Stress conditions such as pathogenic infections, hypoxia and protein misfolding impact protein homeostasis activating a number of stress signaling pathways such as the unfolded protein response (UPR) at the ER and the heat shock response in the cytoplasm along with translational remodeling. We combine state-of-the-art structural tools including functionalized microscopy grids, cryo-electron microscopy (cryoEM) and cryo-electron tomography (cryoET) with biochemical, sequencing, and computational methodologies to obtain molecular details of these stress signaling pathways. By mapping how these systems function, we aim to develop next generation therapeutics combining structure-guided drug design with computational modeling for a range of diseases, including neurodegenerative disorders, cancer, and metabolic diseases where protein quality control is dysfunctional.
For more information view Dr. Sangwan's faculty profile.
Publications
Please see Dr. Sangwan's publications on PubMed.
Contact Information
Contact Dr. Sangwan

Vijay Sarthy
Professor Emeritus, Ophthalmology
Gene regulation, development and functional organization of the vertebrate retina

Vijay Sarthy
Research Area(s): Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology
The pattern of gene expression in eukaryotic cells is strongly influenced by interactions with neighboring cells. When cell-cell interactions are perturbed, changes in cellular gene activity are often observed. In the vertebrate retina, inherited or acquired rod and cone degeneration results in disruption of normal interactions between photoreceptors and their support cells, the Müller cells. Under these conditions many genes such as the glial intermediate filament protein (GFAP) gene, ciliary neurotrophic factor (CNTF) gene and basic fibroblast growth factor (bFGF) gene are upregulated in neighboring Müller cells.
We use techniques such as single cell RT-PCR and differential display to study changes in gene expression patterns in Müller cells. Major goals of our current research are to elucidate the molecular mechanisms responsible for transcriptional activation and to determine the extracellular inductive signal and the signal transduction pathways involved. Our recent cell transfection studies and experiments with GFAP-lacZ transgenic mice suggest that GFAP gene activation in Müller cells is regulated by a cell type-specific, inducible enhancer and that GFAP gene is activated through the JAK-STAT pathway. The work on gene regulation is crucial for development of strategies for using Müller cell-specific promoters to test the biological effects of growth factors and cytokines in animals models of retinal degeneration and more importantly for designing cell type-specific vectors for targeted delivery in gene therapy.
A second project is concerned with molecular cloning, regulation and function of neurotransmitter transporters—a family of membrane proteins that are involved in the uptake of neurotransmitters. We are particularly interested in the role of taurine and glutamate transporters in retinal ischemia and glutamate neurotoxicity. We have already cloned and characterized GABA, taurine and glutamate transporters from retina. We have also localized the transporters to specific retinal cell types and shown that phosphorylation may play a key role in regulating transporter function.
For more information visit Dr. Sarthy's faculty profile page.
Publications
View Dr. Sarthy's publications at PubMed
Contact
Phone 312- 503-3031

Karla Fullner Satchell
Professor, Microbiology-Immunology
Role of bacterial protein toxins in the pathogenesis of Vibrio vulnificus and Vibrio cholerae

Karla Fullner Satchell
Research Area(s): Biochemistry and Structural Biology; Microbiology
My research focuses on the role of secreted protein toxins on bacterial pathogenesis. The toxins we study are members of the MARTX family and are produced by Vibrio cholerae, a pathogen important for the diarrheal disease cholera, and Vibrio vulnificus, a pathogen that causes septicemia and necrotizing fasciitis from seafood consumption as well as wound infections. Our group studies the mechanism of action of these toxins using a combination of cell biology, biochemistry, and structural biology. In addition, we investigate the role of these toxins in pathogenesis using animal and tissue culture models with focus on mechanisms of tissue damage and evasion of innate immune clearance.
For lab information and more, see Dr. Satchell's faculty profile.
Publications
See Dr. Satchell's publications on PubMed
Contact

Jeffrey N Savas
Associate Professor, Neurology (Behavioral Neurology), Pharmacology, Medicine (Nephrology and Hypertension)
Proteins and proteomes responsible for neurodevelopmental and neurodegenerative diseases

Jeffrey N Savas
Research Area(s): Biochemistry and Structural Biology; Neurobiology
We use biochemistry with discovery-based mass spectrometry to identify the protein perturbations which drive synaptopathies and proteinopathies. Groups of perturbed proteins serve as pathway beacons which we subsequently characterizes in hopes of finding new pathogenic mechanisms and potential future therapeutic targets.
For more information view the faculty profile of Jeffrey Savas, PhD or the Savas Lab website.
Publications
Please see Dr. Savas' publications on PubMed.
Contact Information
Jeffrey N Savas, PhD
Assistant Professor in Neurology
312-503-3089

Richard C Scarpulla
Professor, Cell and Developmental Biology
Molecular interactions and physiological functions of transcriptional activators and co-activators involved in the nuclear control of the respiratory apparatus

Richard C Scarpulla
Research Area(s): Cell Biology; Genetics, Epigenetics and Genomics
Our long-term objectives are to further define the molecular interactions and physiological functions of transcriptional activators and co-activators involved in the nuclear control of the respiratory apparatus. Current work in the lab combines molecular and biochemical approaches with the development of cellular and transgenic models to understand in vivo regulatory pathways and mechanisms.
For lab information see Dr. Scarpulla's faculty profile.
Publications
See Dr. Scarpulla's publications on PubMed
Contact
Contact Dr. Scarpulla at 312-503-2946.

Robert P Schleimer
Professor Emeritus, Medicine (Allergy and Immunology)
Molecular and cellular mechanisms underlying pathogenesis of Chronic Rhinosinusitis and actions of anti-inflammatory steroids

Robert P Schleimer
Research Area(s): Immunology; Pharmacology
Our group studies Chronic Rhinosinusitis, a disease of the nose and sinuses that affects over 30 million Americans. Using tissue samples collected from patients undergoing surgery for Chronic Rhinosinusitis through our close collaboration with Clinical Allergists and ENT surgeons, we study the gene and protein expression profiles as well as resident immune cell populations that are altered in disease in order to identify and explore potential mechanisms of disease progression and identify targets for therapeutic development. This work has provided evidence of dysregulation of epithelial immune barrier function, innate immunity, fibrin deposition and adaptive immune responses as potential mechanisms of disease progression. In the realm of epithelial barrier, we are currently investigating the mechanism by which Oncostatin M drives epithelial barrier degeneration. Additional research in our lab is pursuing mechanisms of fibrin deposition and the contributions of alterations in the fibrin system to polyp formation. Other experiments are working to reveal the potential impact of abnormal B cell activation in polyps. Finally, we continue our long running exploration of the cellular and molecular mechanisms of anti-inflammatory actions of glucocorticoid steroids and the causes of apparent steroid resistance in selected individuals and diseases.
For more information, visit the faculty profile of Robert Schleimer, PhD.
Publications
View Dr. Schleimer's publications at PubMed
Contact
Contact Dr. Schleimer at 312-503-0076

Paul T Schumacker
Professor, Pediatrics (Neonatology), Cell and Developmental Biology, Medicine (Pulmonary and Critical Care)
Oxygen sensing in embryonic development, tissue responses to hypoxia and tumor angiogenesis

Paul T Schumacker
Research Area(s): Cancer Biology; Cell Biology; Lung Biology
Our lab is interested in the molecular mechanisms of oxygen sensing and the importance of this process for embryonic development, tissue responses to hypoxia and tumor angiogenesis. We are testing the hypothesis that the mitochondria play a central role in detecting cellular oxygenation and signal the onset of hypoxia by releasing reactive oxygen species (ROS). These signals trigger downstream signal transduction pathways responsible for the transcriptional and post-translational responses of the cell. Transcriptional activation of genes by Hypoxia-Inducible Factor-1 confers protection against more severe hypoxia by augmenting the expression of glycolytic enzymes, membrane glucose transporters and other genes that tend to augment tissue oxygen supply by increasing the release of vascular growth factors such as VEGF, erythropoietin and vasoactive molecules that augment local blood flow. Current experiments are aimed at improving our understanding of how oxygen interacts with the mitochondrial electron transport chain to amplify ROS production and clarifying the targets that they act on to stabilize HIF and activate transcription.
In specific tissues, oxygen sensing is essential for normal function, but it can also contribute to disease pathogenesis. For example, during mammalian development, the lung tissue is hypoxic and blood flow is restricted in the pulmonary circulation in order to prevent escape of oxygen from the pulmonary capillaries to amniotic fluid. At birth, inflation of the lung with air causes an increase in lung oxygen levels, which triggers relaxation of pulmonary arteries. In Persistent Pulmonary Hypertension of the Newborn, failure of the pulmonary circulation to dilate results in elevated pulmonary arterial pressures and significant lung gas exchange dysfunction. We are testing the hypothesis that pulmonary vascular cells sense O2 at the mitochondria and that ROS released from those organelles trigger an increase in cytosolic calcium, which causes smooth muscle cell contraction. In adult patients with hypoxic lung disease, similar activation of hypoxic vasoconstriction can lead to chronic pulmonary hypertension, which can progress to right heart failure. A fuller understanding of the mechanisms of oxygen sensing in health and disease may lead to insights into therapeutic inhibition of this response in disease states.
In solid tumors, consumption of oxygen by highly metabolic tumor cells leads to hypoxia and threatens glucose supplies. Hypoxic tumor cells retain their oxygen sensing capacity and turn on expression of HIF-dependent genes, leading to tumor angiogenesis and increased blood supply, which permits further growth. We are currently exploring the hypothesis that the mitochondrial oxygen sensor is required for this response using pursuing genetic models. A better understanding of how tumor cells detect hypoxia could lead to the discovery of therapeutic approaches that would prevent detection of hypoxia and thereby prevent tumor progression.
For more information visit Dr. Schumacker's faculty profile page
Publications
View Dr. Schumacker's publications at PubMed
Contact
Phone 312-503-1476

Steven J Schwulst
Associate Professor, Surgery (Trauma and Critical Care)
Monocyte and microglia interaction in the etiology and evolution of traumatic brain injury-induced neurodegeneration

Steven J Schwulst
Research Area(s): Computational Biology and Bioinformatics; Immunology; Microbiology; Neurobiology
Dr. Schwulst is an Associate Professor of Surgery at Northwestern University and an attending Trauma and Critical Care surgeon at Northwestern Memorial Hospital. His primary research interests are in post-injury immune dysfunction, chronic neurodegenerative disease (such as Alzheimer's disease and Chronic Traumatic Encephalopathy), and inflammatory bowel disease-associated psychiatric disease all with a particular emphasis on microglia biology, gut-brain axis crosstalk, and gut microbiome. These investigations are carried out in rodent models of traumatic brain injury (TBI) and inflammatory bowel disease (IBD) as well as with fresh human tissue and the generation of humanized rodent models. Current projects include: 1) the role of the post-injury gut microbial structure in neurocognitive/degenerative outcomes after TBI, 2) the role of microglial activation in the etiology and evolution of chronic neurodegeneration after TBI, 3) identifying common molecular pathways between TBI and neuropsychiatric disease and 4) interactions between the IBD-associated gut microbiome, microglia activation, and neuropsychiatric outcomes in inflammatory bowel disease.
For more information, please see Dr. Schwulst's faculty profile.
Publications
See Dr. Schwulst's publications on PubMed
Contact

Patrick C Seed
Professor, Pediatrics (Infectious Diseases), Microbiology-Immunology
Microbial/microbiome genetics, physiology, and pathogenesis to develop diagnostics and treatments for a broad range of pediatric diseases

Patrick C Seed
Research Area(s): Microbiology
The Seed Research Group broadly studies the interactions of mammals and microbes to understand how microbial communities alter health and disease and how pathogenic microbes emerge from those communities to cause specific infections of the urinary tract, blood, and central nervous system. Our work combines clinical human studies, conventional and germ-free animal models, molecular genetics, immunology, biochemistry, structural biology, high-dimensional data computational analysis, and a number of complementary 'omics technologies including genomics, metagenomics, transcriptomics, metatranscriptomics, and metabolomics to understand complex systems. We use the lessons learned from these molecular studies to design new diagnostics and therapeutics.
For lab information and more, see Dr. Seed's faculty profile.
Publications
See Dr. Seed's publications on PubMed.
Contact
Email Dr. Seed
Phone at 312-227-4080

Ali Shilatifard
Professor, Biochemistry and Molecular Genetics
Professor, Weinberg College of Arts and Sciences
Molecular machinery for histone modifications in yeast, Drosophila and human cells

Ali Shilatifard
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics
Chromosomal rearrangements resulting in alterations of gene expression are a major cause of hematological malignancies. Our goal is to advance the understanding of the biochemical and molecular mechanisms of rearrangement-based leukemia and to provide insights into how translocations affect cellular division by altering gene expression. Using mammalian model systems such as tissue culture and mouse genetics, we plan to explore the regulation of gene expression via the MLL gene and its translocation partners found in human leukemia. We are currently defining the molecular composition of the MLL complexes and how translocations alter its biochemical function and integrity, resulting in leukemic pathogenesis. We are also planning to define the mechanism of the targeting of the MLL complex and its histone methyltransferase activity to chromatin to determine its normal cellular functions and its mistargeting and dysregulation in leukemogenesis.
One fusion partner of MLL in acute myelogenous leukemia (AML) is the ELL protein. We show that human ELL functions as a transcription elongation factor. We have identified the Drosophila homolog of ELL and demonstrate it to be essential for development. Drosophila ELL associates with elongating RNA polymerase II in vivo on chromosomes and is a regulator of the Notch signaling pathway. This has suggested to us that human ELL might also participate in the same process.
For lab information and more, see Dr. Shilatifard's faculty profile or visit the Shilatifard Laboratory site.
Publications
View Dr. Shilatifard's publications on PubMed.
Contact
Phone 312-503-5223

Vipul Shukla
Assistant Professor, Cell and Developmental Biology, Medicine (Hematology and Oncology)
Deciphering alternative DNA codes in normal and cancer genomes

Vipul Shukla
Research Area(s): Cancer Biology; Genetics, Epigenetics and Genomics; Immunology; Metabolism and Endocrinology
Our lab applies state-of-the-art genetics, genomics, molecular biology and cell biology techniques to decipher the functions of cytosine modifications and structural conformations as alternative DNA codes in the genome. Decades of research have established how specific DNA sequences control genomic states associated with transcription, chromatin modifications and topological compartmentalization. However, besides helical, linear sequences, the DNA in the genome commonly adopts unusual, non-helical structural conformations and we want to understand the significance of these alternative structural conformations in normal cellular physiology and associated pathologies. As first-steps towards understanding the functions of alternative DNA structures, our lab is studying their abundance in normal and cancer genomes, epigenetic mechanisms regulating their localization and dynamics, and cellular pathways controlled by these structures. These studies have broad implications on many established paradigms in genome biology and will address fundamental questions related to origins of several different cancers with the ultimate goal of identifying vulnerabilities that could be therapeutically targeted.
Our lab also holds strong interest in understanding basic molecular mechanisms regulating immune responses. We are particularly interested in understanding how changes in the metabolic outputs, that are associated with distinct stages of B (and T) cell differentiation impacts their epigenetic landscapes. We aim to uncover these mechanisms with the ultimate objective to design approaches by which we could engineer the desired epigenetic states in immune cells to enhance the fidelity of immune responses.
For lab information and more, see Dr. Shukla's faculty profile or visit the Shukla Laboratory site.
Publications
View Dr. Shukla's publications on PubMed.
Contact

Benjamin D Singer
Associate Professor, Medicine (Pulmonary and Critical Care), Biochemistry and Molecular Genetics
Exploring respiratory failure

Benjamin D Singer
Research Area(s): Genetics, Epigenetics and Genomics; Immunology; Lung Biology; Metabolism and Endocrinology

Gregory A Smith
Professor, Microbiology-Immunology
Cell and molecular biology of herpesvirus invasion of the nervous system

Gregory A Smith
Research Area(s): Cell Biology; Developmental and Stem Cell Biology; Immunology; Microbiology; Neurobiology
We investigate the relationship between infection of the nervous system by herpesviruses and disease outcome. Some of the most traumatic diseases – including polio, rabies and encephalitis – result from infections of the nervous system. In contrast, herpesviruses are highly proficient at infecting the nervous system, yet normally do not cause neurological disease. This is achieved in part by self-imposed restrictions encoded within the viruses that limit viral reproduction and prevent dissemination into the brain. For the individual, this results in a relatively benign infection, yet the virus becomes a life-long occupant of the nervous system that will periodically reemerge at body surfaces to infect others. Unfortunately, this infectious cycle can go awry resulting in several forms of severe disease (i.e. keratitis and encephalitis).
We have pioneered methods to genetically manipulate herpesviruses and visualize individual viruses in living neurons. Using these methods, we are studying the mechanisms by which the virus achieves its stringently controlled infectious cycle. Current genetic manipulations are based on a full-length infectious clone of the herpesvirus genome. The clone was made as a bacterial artificial chromosome (BAC) in E. coli. Transfection of purified E. coli BAC plasmid into permissive eukaryotic cells is sufficient to initiate viral infection, allowing for immediate examination of viral mutant phenotypes in a variety of biological assays. For example, by fusing the green fluorescent protein (GFP) to a structural component of the viral capsid, individual viral particles can be tracked within the axons of living neurons during both entry and egress phases of the infectious cycle. Studies in culture can be complemented by examining the pathogenesis of mutant viruses in rodent models of infection.
Using these methods, we have discovered key aspects of cellular infection, viral assembly and intracellular transport. Looking forward, we are continuing to pursue our multidisciplinary approach of combining neuroscience, cell biology, bacterial genetics and virology to better understand these important pathogens.
For lab information and more, see Dr. Smith's faculty profile.
Publications
See Dr. Smith's publications on PubMed.
Contact

Richard S Smith
Assistant Professor, Pharmacology, Pediatrics
Ion channels in human brain development and brain malformation diseases

Richard S Smith
Research Area(s): Biochemistry and Structural Biology; Developmental and Stem Cell Biology; Neurobiology
Dr. Smith’s research examines ion channel contributions to human brain development and brain malformation diseases. Since 2016 Dr. Smith has received continuous funding support from the National Institute of Health to evaluate the cellular, biophysical, and genetic mechanisms underlying aberrant neurophysiology related to developmental channelopathies.
Current Projects
Ion channel diseases affecting brain development.
Evaluating children with cerebral cortex malformation diseases, we identified a role for a sodium channel (SCN3A), and a Na+/K+-ATPase (ATP1A3) in the developing human cortex. This developmental channelopathy affects several populations of cells in the fetal human cortex, which at critical times of development results in atypical neocortical formation.
Bioelectricity in human brain development and evolution.
We use animal model systems (i.e. mouse and ferret) to unravel the non-canonical roles of ion channels in generating “Bioelectric” patterns of neuronal activity across mammalian brain development. In conjunction with Human Genetics studies, these studies enable dissection of species-specific neurophysiology, identifying key cellular and biophysical mechanisms unique to the human brain and disease pathology.
Human induced pluripotent stem cell (iPSCs) screening and neuronal disease modeling platform.
Using human iPSCs, we model various ion channel diseases found in human patients. In these neuronal culture systems, we use both patch-clamp and high-throughput neurophysiology assays to evaluate pharmacology and precision therapeutics.
For more information, see Dr. Smith's faculty profile.
Publications
See Dr. Smith's publications on PubMed.
Contact
Contact Dr. Smith.

Wilton Snead
Assistant Professor, Cell and Developmental Biology
Decoding the biophysical mechanisms of RNA localization

Wilton Snead
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Genetics, Epigenetics and Genomics
All cells, from bacteria to neurons, depend on the precise localization of messenger RNA (mRNA) transcripts at subcellular locations to control local protein production. mRNA localization is mediated primarily by RNA-binding proteins, which recognize mRNA targets and bring them to their destinations. mRNAs contain many kinds of information that control interactions with RNA-binding proteins, including “zip code” sequence motifs, secondary or tertiary structural elements, and post-transcriptional nucleotide modifications. These features also determine interactions with biomolecular condensates, which are membrane-less subcellular compartments comprising dense collections of interacting molecules. Although all of the mRNA features listed above contribute to localization, it remains unclear how RNA-binding proteins and biomolecular condensates decode this complex information to escort mRNAs to their sites of function.
While we know some of the biomolecular “parts lists” that mediate mRNA localization, we are left with the challenging task of figuring out how these parts fit together. In the Snead lab, we adopt a reverse-engineering approach to this challenge: by reassembling biological systems from component parts, we can understand how molecules recognize each other and how their interactions are perturbed in disease. Our lab leverages quantitative microscopy, live-cell imaging, and bottom-up biochemical reconstitution to understand the biophysical rules that control mRNA localization in cells. Current projects include (1) understanding how a condensate controls mRNA retention in the nucleus, (2) uncovering how dysregulated mRNA nuclear retention impacts cancer cell development and motility, (3) deciphering how endomembrane-bound RBPs sort mRNA cargos for long-distance transport, and (4) elucidating how membrane-localized mRNAs are translated into proteins.
To learn more, please visit Dr. Snead’s faculty profile and lab website.
Publications
See the Snead lab’s publications on the Snead lab website.
Contact
Contact Dr. Snead.

Beatriz Sosa-Pineda
Professor, Medicine (Nephrology and Hypertension)
Regulation of acinar cell development and plasticity in the pancreas, hepatic cell fate, and liver zonation as well as mechanisms that promote pancreas metastasis

Beatriz Sosa-Pineda
Research Area(s): Cancer Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics
Using genetically modified mouse models and cutting-edge technologies, we investigate how the complex architecture of the mammalian pancreas and liver is established during development. We also investigate how acute or chronic injury affect liver zonation and exocrine pancreas homeostasis, and the role of chromosomal instability in pancreatic tumor formation and metastasis.
For more information, visit the faculty profile of Beatriz Sosa-Pineda, PhD or the Sosa-Pineda lab web site.
Publications
View Dr. Sosa-Pineda's publications at PubMed
Contact
Phone 312-503-2296

Thomas Stoeger
Assistant Professor, Medicine (Pulmonary and Critical Care)
Understanding and mitigating aging and age-related diseases

Thomas Stoeger
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Lung Biology
We are interested in understanding and mitigating aging and age-related diseases through massive data-mining and machine learning applied to the entire biomedical literature. This approach helps us identify strong biological associations beyond the usual search heuristics employed in most laboratories, allowing us to explore previously unexamined areas in greater detail.
Using this data-driven strategy, we've successfully pinpointed the primary cause of transcriptional changes occurring during aging. We are currently expanding our research to investigate epigenetic and proteomic changes, as well as conditions that either accelerate or delay aging.
Furthermore, our work has led to collaborations, particularly with Northwestern's SCRIPT systems biology center, focused on pneumonia. Additionally, we are in the process of preparing the publication of the largest existing knowledge base about human genes, and, in partnership with our collaborators at NIH, we are pioneering computational AI studies on the contemporary history of science.
For lab information and more, see Dr. Stoeger's faculty profile or lab website.
Publications
See Dr. Stoeger's publications on PubMed.
Contact
Contact Dr. Stoeger

Ronen Sumagin
Associate Professor, Pathology (Experimental Pathology)
Contributions of immune cell-mediated inflammation to development and progression of colorectal cancers

Ronen Sumagin
Research Area(s): Cancer Biology; Cell Biology; Immunology
Immune cells are critical for host defense, however immune cell infiltration of mucosal surfaces under the conditions of inflammation leads to significant alteration of the tissue homeostasis. This includes restructuring of the extracellular matrix and alterations in cell-to-cell adhesions. Particularly, immune cell-mediated disruption of junctional adhesion complexes, which otherwise regulate epithelial cell polarity, migration, proliferation and differentiation can facilitate both tumorigenesis and cancer metastasis. Our research thus focuses on understanding the mechanisms governing leukocyte induced tissue injury and disruption of epithelial integrity as potential risk factors for tumor formation, growth and tissue dissemination.
For publication information, please see Dr. Sumagin's faculty profile page or laboratory site.
Publications
See Dr. Sumagin's publication in PubMed
Contact information
Ronen Sumagin, PhD
Assistant Professor in Pathology
312-503-8144

Geoffrey T Swanson
Professor, Pharmacology
Professor, Weinberg College of Arts and Sciences
Glutamate receptors in the modulation of neurotransmission and induction of synaptic plasticity

Geoffrey T Swanson
Research Area(s): Neurobiology; Pharmacology
Geoffrey Swanson’s, PhD, laboratory studies the molecular and physiological properties of receptor proteins that underlie excitatory synaptic transmission in the mammalian brain. Current research focuses primarily on understanding the roles of kainate receptors, a family of glutamate receptors whose diverse physiological functions include modulation of neurotransmission and induction of synaptic plasticity. We are also interested in exploring how kainate receptors might contribute to pathological processes such as epilepsy and pain. The laboratory investigates kainate receptor function using a diverse group of techniques that include patch-clamp electrophysiology, selective pharmacological compounds, molecular and cellular techniques and gene-targeted mice.
Current Projects
- Isolation and characterization of new marine-derived compounds that target glutamate receptors
- Kainate receptors in hippocampal synaptic transmission
- Mechanisms of kainate receptor assembly and trafficking
For lab information and more, see Dr. Swanson’s faculty profile and lab website.
Publications
See Dr. Swanson's publications on PubMed.
Contact
Contact Dr. Swanson at 312-503-1052.

Jacob I Sznajder
Professor, Medicine (Pulmonary and Critical Care), Cell and Developmental Biology
Mechanisms of acute lung injury as related to aging, high CO2, low oxygen, lung cancer and influenza infection

Jacob I Sznajder
Research Area(s): Cell Biology; Lung Biology; Microbiology; Pharmacology
Seasonal influenza infection affects a significant proportion of the population in the US and worldwide and while most patients infected with influenza A virus (IAV) recover without sequelae, in many patients influenza virus infection may cause ARDS. Alveolar epithelial cells (AEC) are targets for IAV and play an important role in mounting the initial host response. The Sznajder Lab hypothesizes that the alveolar epithelium plays an important effector role in protecting the lung from severe injury. Findings indicate that the degradation of PKCζ, which triggers the down-regulation of Na,K-ATPase, by the E3 ligase HOIL-1L decreases AEC death. HOIL-1L is a member of the Linear Ubiquitination Assembly Complex (LUBAC) and the lab studies whether LUBAC participates in the modulation of the inflammatory intensity in the lung epithelium during IAV infection. Also, they are investigating the mechanisms by which modest inhibition of the Na,K-ATPase, whether pharmacologic inhibition of the Na,K-ATPase by cardiotonic steroids such as ouabain are protective by inhibiting virus replication.
Studies suggest that signals from the injured lung during IAV infection disrupt skeletal muscle proteostasis and contribute to skeletal muscle dysfunction. The slower recovery of the skeletal muscle function in aged mice during IAV pneumonia is the consequence of diminished proteostatic reserve in cells responsible for regenerating the damaged skeletal muscle.
Hypercapnia (high pCO2) is observed in patients with lung diseases such as chronic obstructive pulmonary disease (COPD), broncho-pulmonary dysplasia and advanced neuromuscular diseases. The lab hypothesizes that hypercapnia promotes the ubiquitin-proteasome mediated muscle degradation and impairs the function of muscle satellite cells required for its regeneration.
Alveolar fluid reabsorption is effected by vectorial Na+ transport via apical Na+ channels and basolateral Na,K-ATPase of the alveolar epithelium. We and others have reported that β-adrenergic agonists upregulate the Na,K-ATPase in AEC by increasing the traffic and recruitment of Na,K-ATPase containing vesicles into the cell membrane, resulting in increased catalytic activity. Moreover, GPCR-mediated upregulation of the Na,K-ATPase resulted in increased alveolar fluid clearance in normal lungs and in rodent models of lung injury. We are investigating mechanisms of Na,K-ATPase regulation and active Na+ transport in lungs which will help with the design of new strategies to increase lung edema clearance.
For more information visit the faculty profile of Jacob Sznajder, MD.
Publications
View Dr. Sznajder's publications on PubMed
Contact
Contact Dr. Sznajder at 312-908-7737 or the Sznajder Lab at 312-503-1685.

Colby S Thaxton
Associate Professor, Urology
Fabricating new nanomaterials and translational nanotechnology with regard to nanoparticle-based molecular diagnostics and nanotherapeutics

Colby S Thaxton
Research Area(s): Biochemistry and Structural Biology; Pharmacology
Shad Thaxton MD, PhD, invented, synthesized, and characterized the first biomimetic high-density lipoprotein nanoparticles. High density lipoproteins (HDLs) are natural nanoparticles that circulate in the human body to carry cholesterol. Cholesterol carried by HDLs is often referred to as “good cholesterol” because HDL blood levels inversely correlate with the development of cardiovascular disease. The Thaxton Group utilizes a gold nanoparticle scaffold to assemble the natural surface chemical components of HDLs to create synthetic HDL nanoparticles. The HDL nanoparticles recapitulate the size, shape, surface chemistry, and cholesterol binding properties of natural HDLs. As such, The Thaxton Lab is using these unique biomimetic materials to better understand the structure-function properties of natural HDLs and also as potential therapies for atherosclerosis and heart disease.
For more information, see the faculty profile of C. Shad Thaxton, MD, PhD
Publications
View Dr. Thaxton's other publications at PubMed
Contact
Phone 312-503-1826

Benjamin Thomson
Assistant Professor, Ophthalmology
Links between endothelial function and vision

Benjamin Thomson
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Developmental and Stem Cell Biology
Endothelial dysfunction is a major cause of vision loss, playing a key role in diseases including age-related macular degeneration, diabetic retinopathy and glaucoma. Using mouse genetics, animal disease models and a combination of single cell RNA-sequencing and histological approaches, our lab is focused on understanding the role of the vasculature in these diseases, including glaucoma and age related macular degeneration. By elucidating molecular connections between endothelial dysfunction and vision loss, we aim to identify novel therapeutic targets and translate these discoveries into patient care.
While endothelial dysfunction is a component of many eye diseases, the importance of ocular vasculature in age related macular degeneration is widely understood and is the basis for the life-altering anti-VEGF therapies that target choroidal neovascularization associated with these conditions. The choroid and choriocapillaris (CC) form a unique vascular bed in the back of the eye that is vital for maintenance of the retinal photoreceptors and retinal pigment epithelium (RPE). In addition to the well-described link with AMD, choroidal dysfunction is tied to the poorly understood spectrum of pachychoroid diseases including polypoidal choroidal vasculopathy (PCV), which can lead to irreversible loss of vision. Despite their clinical impact, little is known about pathogenesis or optimal treatment of PCV and other pachychoroid diseases, or why some patients with defects in the choroidal vasculature develop geographic atrophy or neovascular AMD and others PCV. Ongoing research in our lab seeks to answer this question, using animal models, single cell RNA sequencing and in vivo imaging to gain mechanistic insights into pachychoroid biology, understand mechanisms by which choriocapillaris attenuation lead to choroidal dysfunction, and identify genes and pathways which can be targeted for future therapies.
For additional information, visit the faculty profile of Dr. Thomson
Publications
View Dr. Thomson's publications at PubMed
Contact
Contact Dr. Thomson

Edward B. Thorp
Professor, Pathology (Experimental Pathology)
Molecular mechanisms by which the immune system regulates wound repair, inflammation resolution, and tissue regeneration

Edward B. Thorp
Research Area(s): Biochemistry and Structural Biology; Cardiovascular and Renal Biology; Cell Biology; Genetics, Epigenetics and Genomics; Immunology; Metabolism and Endocrinology; Microbiology; Pharmacology
The Edward Thorp Lab studies the crosstalk between immune cells and the cardiovascular system and, in particular, within tissues characterized by low oxygen tension or associated with dyslipidemia, such as during myocardial infarction. In vivo, the lab interrogates the function of innate immune cell phagocytes, including macrophages, as they interact with other resident parenchymal cells during tissue repair and regeneration. Within the phagocyte, the influence of hypoxia and inflammation on intercellular and intracellular signaling networks and phagocyte function are studied in molecular detail. Taken together, our approach seeks to discover and link basic molecular and physiological networks that causally regulate disease progression and in turn are amenable to strategies for the amelioration of cardiovascular disease.
For additional information, visit the Thorp Lab site or view the faculty profile of Edward B Thorp, PhD.
Publications
View Dr. Thorp's publications at PubMed
Contact
Contact Dr. Thorp or the Thorp lab at 312-503-3140.

Praveen Thumbikat
Professor, Urology, Pathology
Benign prostate diseases, chronic prostatitis/chronic pelvic pain syndrome

Praveen Thumbikat
Research Area(s): Immunology; Reproductive Biology
The focus of research in the laboratory is to understand the pathogenesis of genitourinary diseases with emphasis on benign prostate disease in humans. Inflammation is a significant finding in a variety prostate diseases including prostatitis, BPH and prostate cancer. We study microbial and autoimmune mediated inflammation and innate and adaptive immune mechanisms in prostate disease. A particular area of interest is chronic prostatitis/chronic pelvic pain syndrome (CP/CPPS), a debilitating medical condition characterized by dysuria and pain. Projects in the lab use a combination of in vitro studies, animal models and clinical specimen assays to examine questions of interest such as the role of chemokines and T-cells in chronic pelvic pain.
For more information, see the faculty profile of Praveen Thumbikat, PhD.
Publications
View Dr. Thumbikat's publications at PubMed
Contact
Phone 312-503-1050

Iris Titos Vivancos
Assistant Professor, Biochemistry and Molecular Genetics, Psychiatry and Behavioral Sciences
Peripheral organ regulation of behavior

Iris Titos Vivancos
Research Area(s): Cell Biology; Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Neurobiology
Research aiming to understand how behavior arises has historically focused on the brain. However, the brain is not isolated from the rest of the body: peripheral organs like the gut produce a repertoire of unexplored signals that communicate with the brain and influence behavioral decisions. The focus of the lab is to identify novel peripheral organ signaling pathways and determine how they influence behaviors such as sleep and substance use disorder. Sleep is essential for life, and poor sleep quality represents a risk factor for metabolic and psychiatric disorders. While sleep duration has been extensively investigated, sleep depth, which is equally important because of its restorative functionovers, is understudied due to the challenges in measuring it. We have developed a behavioral setup that allows us to investigate how dietary manipulations and their effect on gut secretory signaling pathways affect sleep depth. Diet imbalances are also a strong predictor for substance use disorder and we have developed novel assays showing that dietary changes can alter psychostimulant preference. Our overarching goal is to provide crucial and missing molecular mechanistic understanding of how inter-organ signals shape essential and maladaptive behaviors, to enable novel preventive and therapeutic strategies.
For additional information, visit the faculty profile of Dr. Titos Vivancos
Publications
View Dr. Thomson's publications at PubMed
Contact
Contact Dr. Titos Vivancos
Sergey M Troyanovsky
Professor, Dermatology, Cell and Developmental Biology
Mechanisms of adherens junctions assembly, dynamics and signaling
Sergey M Troyanovsky
Research Area(s): Cell Biology
The Troyanovsky lab’s research focuses on cadherin, intercellular adhesion and signaling. Classic cadherins are critical proteins mediating cell-cell adhesion and various signaling pathways responsible for cellular proliferation, differentiation and morphogenesis. Abnormalities in this system are causal factors in many pathologies, including cancer. The molecular mechanisms of cadherin-based adhesion, however, are largely unknown. How do cadherins establish the adhesion contact? How do they interact with the cytoskeleton? What are the signaling pathways they control? Our laboratory's work is centered around these questions. We are currently working on the following specific projects:
- An individual cadherin molecule’s adhesion site is very weak. To mediate tight adhesion, cadherin molecules form clusters. Recently our lab showed that cadherin clustering is based on two different mechanisms. First, using an extracellular cis-binding site, cadherin sticks laterally in small groups. Additional clustering is promoted by the actin cytoskeleton, binding to which limits cadherin diffusion. The aim of our current study is to understand the regulation of cadherin clustering through modulation of the cadherin-actin filament coupling.
- The formation of cadherin adhesive clusters interconnected to the cytoskeleton is not sufficient to establish functional intercellular junctions. The junctions stimulate formation of actin bundles that is required for epithelial cells to organize their actin cytoskeleton. How adherens junctions initiate actin bundle formation is another direction in our research.
- Cadherin is not the only transmembrane protein in adherens junctions. These structures contain adhesion proteins from the nectin family as well as numerous signaling proteins. We showed that one of such proteins, gamma-secretase, interacts with E-cadherin through p120-catenin. The roles of nectins and gamma-secretase and the ways they are recruited into adherens junctions are also areas of focus in our lab.
For more information see Sergey Troyanovsky’s, PhD, faculty profile.
Publications
View Dr. Troyanovsky's publications at PubMed.
Contact
Phone the Troyanovsky Lab at 312-503-9275

Margrit Urbanek
Associate Professor, Medicine (Endocrinology, Metabolism and Molecular Medicine)
Susceptibility genes for complex diseases

Margrit Urbanek
Research Area(s): Genetics, Epigenetics and Genomics; Metabolism and Endocrinology; Reproductive Biology
Dr. Urbanek’s research focuses on the identification of susceptibility genes for complex diseases. Her approach to this research is to use family-based gene-mapping techniques and population-based association studies in conjunction with molecular techniques to identify and verify genes and pathways contributing to the pathogenesis of genetically complex diseases. Specifically, she is carrying out studies to identify susceptibility genes for polycystic ovary syndrome (PCOS) that map to Chr19p3.13. She has previously shown that this region shows linkage and association with PCOS in a large set of families. Other projects focus on identifying candidate genes for gestational diabetes and glycemic control during pregnancy and identifying genetic variation contributing to extreme obesity
Research Topics
Identification of sequence variants in PCOS candidate genes
Identification of candidate genes for contributing glycemic control during pregnancy and to gestational diabetes
Genetic variation contributing to extreme obesity
Linkage and family-based association studies
Haplotype analysis
Genome-wide association studies
For more information, visit Dr. Urbanek's faculty profile page.
Publications
View Dr. Urbanek's publications at PubMed.
Contact
Phone 312-503-3658

Robert J Vassar
Professor, Neurology (Behavioral Neurology), Cell and Developmental Biology
Role of Ab and BACE1 in normal biological processes and in disease mechanisms of relevance to Alzheimer’s disease

Robert J Vassar
Research Area(s): Neurobiology
Alzheimer’s disease (AD) is the leading cause of dementia in the elderly. The progressive degeneration of neurons in regions of the brain important for cognition causes the dementia that slowly robs AD patients of their memories, personalities and eventually their lives.
AD pathology is characterized by two microscopic brain lesions:
- Amyloid plaques: Extracellular deposits of the beta-amyloid peptide (Ab), and the longer 42 amino acid form, Ab42, which is strongly associated with autosomal dominant forms of familial AD
- Neurofibrillary tangles
Ab is generated from the amyloid precursor protein (APP) by endoproteolysis from two proteases called the b- and g-secretases. The b-secretase, a novel aspartic protease termed BACE1, was initially cloned and characterized by our group (Vassar, et al., 1999). BACE1 is required for the generation of all forms of Ab, including Ab42, and therefore is a prime drug target for the treatment of AD.
Our ongoing research focuses on the role of Ab and BACE1 in normal biological processes and in disease mechanisms of relevance to AD, including:
- The functions of BACE1 and the homologue, BACE2 and the cell biology of Ab in neurons
- The role of inflammation in AD pathophysiology
- Novel transgenic and knockout mouse models of AD
- Molecular changes that may occur during brain aging leading to neurodegeneration
For lab information and more, visit the Vassar Lab website or see Dr. Vassar's faculty profile.
Publications
See Dr. Vassar's publications on PubMed.
Contact
Phone: 312-503-3361

Douglas E Vaughan
Professor, Medicine (Cardiology)
Plasminogen activator system in cardiovascular disease

Douglas E Vaughan
Research Area(s): Cardiovascular and Renal Biology; Cell Biology
Dr. Vaughan directs a multidisciplinary research group focused on investigating the role of the plasminogen activator system in cardiovascular disease. Active experimental programs are underway at the molecular and cellular level in animals and in humans. Transgenic and knockout mice are used in a variety of studies designed to explore the tissue-specific expression of PAI-1 in vivo and the role of the fibrinolytic system in vascular disease and tissue remodeling.
For more information visit Dr. Vaughan's faculty profile page.
Publications
View Dr. Vaughan's publications at PubMed.
Contact

Lisandra Vila Ellis
Assistant Professor, Cell and Developmental Biology
Endothelial cell heterogeneity in lung development and disease

Lisandra Vila Ellis
Research Area(s): Cardiovascular and Renal Biology; Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Lung Biology
Gas diffusion in the mammalian lung takes place in the alveoli, where a thin epithelium is surrounded by a capillary network. Traditionally, these alveolar capillaries were considered homogeneous until Dr. Vila Ellis' postdoctoral research, which unveiled a novel population of endothelial cells in the mouse lung. Interestingly, the failure to form the alveolar capillary network correctly has been observed in various pulmonary diseases characterized by vascular simplification, such as bronchopulmonary dysplasia (BPD), commonly seen in premature births. Furthermore, recent single-cell RNA sequencing profiling of the human lung by others has also identified a similar endothelial cell population, underscoring the relevance of this research to human diseases.
In the Vila Ellis Lab, our focus lies in investigating this newly discovered heterogeneity during development and disease. Our ultimate objective is to elucidate the transcriptional mechanisms underlying endothelial heterogeneity, endothelial cell fate, and the molecular and cellular components of lung angiogenesis. By doing so, we aim to provide crucial insights into fundamental biology questions.
For more information, see Dr. Vila Ellis's faculty profile or the lab website.
Publications
See publications on PubMed.
Contact
Contact Dr. Vila Ellis

Derek F Walsh
Professor, Microbiology-Immunology
Large DNA Virus Infections

Derek F Walsh
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Genetics, Epigenetics and Genomics; Immunology; Microbiology
The Walsh lab studies three main themes in the area of infection by large DNA viruses:
- Ribosome manipulation and translational control by poxviruses
- Innate immune sensing and evasion during poxvirus infection
- Cellular remodeling during cytomegalovirus infection
For lab information and more, see Dr. Walsh's faculty profile and the lab website.
Publications
See publications on PubMed.
Contact
Contact Dr. Walsh at 312-503-4292

Theresa L Walunas
Associate Professor, Medicine (General Internal Medicine), Medical Social Sciences, Microbiology-Immunology, Preventive Medicine (Biostatistics and Informatics)
Using electronic health record data to improve health outcomes and care for patients with immunological disease, particularly autoimmunity

Theresa L Walunas
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Immunology
Dr. Walunas’s primary research interest is the development and application of novel informatics methods to understand the mechanisms of autoimmune disease. This includes EHR-based phenotyping methods to identify autoimmune disease pathways, identification of environmental, social and behavioral determinants of health in medical records, and use of EHRs to track and improve health outcomes for patients with autoimmune disease.
For more information, visit Dr. Walunas's faculty profile page.
Publications
View Dr. Walunas's program publications at PubMed.
Contact
Walunas Lab: 312-503-3397

Chyung-Ru Wang
Professor, Microbiology-Immunology
Antigen presentation, T-cell development and regulation, infectious diseases and autoimmune diseases

Chyung-Ru Wang
Research Area(s): Immunology
Our lab focuses on two of the MHC class Ib molecules, H2-M3 and CD1. These molecules have unusual binding specificity for antigens that are conserved in bacteria. H2-M3 presents N-formylated peptides to cytotoxic T cells while CD1 presents lipid antigens to several distinct subsets of T cells. The high degree of conservation of these microbial antigens combined with the limited polymorphism of M3 and CD1 make these two molecules attractive targets for T-cell based vaccines against intracellular pathogens for a genetically diverse population. We have generated several animal models to examine the roles of M3 and CD1 in T cell development, autoimmune diseases and defense against infectious agents, including Listeria monocytogenes and Mycobacterium tuberculosis. Using these model systems, we study the mechanisms that regulate the selection and in vivo function of M3-restricted and CD1-restricted T cells. Additionally, these models are used to characterize novel microbial antigens recognized by MHC class Ib-restricted T cells. Studies on this relatively uncharacterized segment of the mammalian immunologic repertoire may lead to improved methods for vaccination against infectious diseases.
For lab information and more, see Dr. Wang's faculty profile.
Publications
See Dr. Wang's publications on PubMed.
Contact
Contact Dr. Wang at 312-503-9748 or the lab at 312-503-1093.

Lu Wang
Assistant Professor, Biochemistry and Molecular Genetics
Mutations in epigenetic factors that contribute to human cancer development

Lu Wang
Research Area(s): Cancer Biology; Pharmacology
Human Cancer Development: Understanding the Important Functions of Epigenetic Factor Mutations
Mutations and/or translocations within genes that encode for epigenetic factors, such as histone protein lysine methyltransferases (KMTs), lysine demethylases (KDMs), and DNA methyltransferases (DNMTs) are all common mechanisms involved in driving tumorigenesis (Cancer Cell. 2019, Feb 11; 35(2):168-176). We utilize state-of-the-art technologies that are designed to conduct epigenetic-related experiments, which allow us to directly uncover the underlying mechanisms of how mutations in epigenetic factors contribute to human cancer development (Nat Med. 2018, Jun; 24(6):758-769).
Novel Cancer Treatment Options: Targeting Dys-Regulated Epigenetic Factors
Misregulation of histone/DNA modifiers have emerged as a common therapeutic target option for treatments of different human diseases, including cancer. (Genes Dev. 2017, Oct 15; 31(20):2056-2066), Cancer Cell. 2014, Jan 13; 25(1):21-36, Sci Adv. 2015 Oct 9; 1(9):e1500463). Currently, several protein methyltransferases and demethylases have been identified, but their physiological significance has just begun to be elucidated. Our goal is to understand the relationship between dys-regulated epigenetic factors and cancer development through the use of these advanced technologies, such as CRISPR screening and experiments involving small inhibitor molecules. As a result, this could lead us to generate potential cancer treatment options by identifying the druggability of selected epigenetic factors, in order to develop a novel and more precise use of a drug that can be translational to clinical applications.
For lab information and more, see Dr. Wang’s faculty profile and lab website.
Publications
See Dr. Wang’s publications.
Contact
Contact Dr. Wang.

Shih-Ting Wang
Assistant Professor, Medicine (Pulmonary and Critical Care)
Assistant Professor, McCormick School of Engineering
Enabling medical technologies and biomaterials

Shih-Ting Wang
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Pharmacology
Well-defined and multifunctional materials are required to probe and perturb complex biological systems and improve the quality of human health. We employ interdisciplinary approaches from materials science, chemistry, and experimental biology to create novel biomaterials with translational potential. These materials are primarily made from natural or synthetic biomolecular building blocks (e.g., protein, nucleic acids) programmed with application-specific sequence, structure, and properties. We are currently applying the materials to sense and intercept cancer, inflammation, and fibrosis. Our materials often work by controlled orchestration of two domains: 1) a “receiving” domain that senses the biochemical signals or bind to target molecules, and 2) a signal transduction domain that was turned on upon signal received, enabling remote sensing from outside of the body or triggering drug delivery. We create these materials by synthesizing the building blocks and engineering to control nanoscale architectures. These materials contain chemical linkages or intrinsic physical properties that can transduce biological signals with high sensitivity and specificity.
For lab information and more, see Dr. Wang’s Feinberg or McCormick faculty profile.
Publications
See Dr. Wang’s publications.
Contact
Contact Dr. Wang.

Daniel Martin Watterson
Professor, Pharmacology
Role of protein phosphorylation pathways in disease onset and progression and their potential as drug discovery targets

Daniel Martin Watterson
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Cell Biology; Immunology; Neurobiology; Pharmacology
Current Projects
The role of calmodulin (CaM) mediated signal transduction pathways in physiology and pathophysiology
- Using of emerging technologies to understand how CaM and a CaM-regulated enzyme could be encoded, expressed, regulated and assembled into a calcium signal transduction complex
- Using of integrative (in vivo) chemical biology and molecular genetics to gain insight into how landmark CaM-regulated protein kinases are involved in physiology and pathophysiology
Integrative chemical biology and development of novel therapeutics for attenuation of disease progression
- Using the “smart chemistry” approach integrated with “smart biology” screens for rapid discovery of novel small molecules with potential use in targeting pathophysiology progression related to diseases ranging from neurological disorders, cancer, inflammatory conditions, cardiovascular and pulmonary disease
- Discovering and developing novel small molecule compounds that selectively attenuate the increased production of proteins called proinflammatory cytokines, which can cause tissue injury and disease when produced in excess
We ultimately hope to find, by targeting pathophysiology mechanisms which contribute to disease progression, a series of novel small molecules with potential to be effective against a variety of disorders.
For lab information and more, see Dr. Watterson’s faculty profile.
Publications
See Dr. Watterson's publications on PubMed
Contact
Contact Dr. Watterson at 312-503-0657.

Samuel E Weinberg
Assistant Professor, Pathology (Experimental Pathology)
Metabolic control of immunity

Samuel E Weinberg
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Immunology; Metabolism and Endocrinology; Microbiology
My laboratory is interested in understanding how changes in immune cell metabolism that occur in diseased environments such as respiratory viral infection or malignancy shape the overall phenotype and function of an immune response. Recent studies, including my previous work, highlight the central role of cell metabolism in dictating immune cell fate and function. These data support an emerging paradigm in which cell metabolism integrates cell-intrinsic and environmental cues to regulate immune cell phenotype and effector function. Furthermore, this model suggests that immunity can be shaped by manipulating immune cell metabolism potentially by employing metabolites or metabolic inhibitors as novel adjuvants.
Despite the potential promise of metabolism-based therapies, our current knowledge of immunometabolism has some critical limitations. For one, the majority of studies evaluating immune cell metabolism have primarily relied on in vitro culture systems that generate immune cells without clear physiologic relevance. In addition, as cell number has historically been a limiting factor for the rigorous assessment of metabolism, little is known about the role of metabolism in critical immune cell types and immunologic processes. Finally, this issue has been further worsened due to a lack of high-quality genetic tools to specifically alter metabolic function in specific rare immune cell types. These limitations have produced a situation where there is an increasing body of immunometabolic knowledge of unclear physiologic and clinical relevance.
The main goal of my group is to unravel how metabolic processes in rare immune cell populations regulate the generation and function adaptive immune responses in clinically relevant contexts. Specifically, the laboratory utilizes a combination of unbiased CRISPR screening, an innovative metabolomics method, high parameter flow cytometry, single cell transcriptomics, and rigorous animal models to identify novel metabolic regulators of immune cell function in a wide range of conditions including cancer, infection and autoimmunity. These differing approaches provide complementary pathways for identifying novel immunometabolic regulators, which can be used to improve vaccination and immunotherapy strategies by utilizing metabolism as a novel modulator of immunity.
For more information, visit the faculty profile of Dr. Weinberg.
Publications
View Dr. Weinberg's publications at PubMed
Contact Us
Contact Dr. Weinberg.

Adam J Williams
Associate Professor, Medicine (Allergy and Immunology)
Molecular mechanisms of adaptive immunity

Adam J Williams
Research Area(s): Genetics, Epigenetics and Genomics; Immunology
The Eisenbarth-Williams lab focuses on defining the cellular and molecular mechanisms that regulate adaptive immunity and antibody responses. The development of these responses relies on the interactions between three immune cells – dendritic cells (DCs), T cells and B cells. In addition, epithelial cells of the lung and intestine are important players in coordinating the immune system. We study how these three cell immune types and epithelial cells communicate to shape different types of immune responses, some protective in the case of vaccination and some harmful in the case of allergy and alloimmunization. By utilizing human samples to guide our studies and mouse models to test new mechanistic paradigms, we have identified novel and unexpected cell subsets and molecular functions. The fundamental principles governing the interaction between DCs, T cells, B cells and the epithelial are the same across these seemingly disparate responses, yet specialization in the subset of each cell type, the specific niche for the interaction and the cellular signals exchanged between the cells dictates what type of immune response is generated. This knowledge can be harnessed to induce protective immune responses and subvert pathogenic ones.
For more information, visit the faculty profile of Dr. Williams.
Publications
View Dr. Williams publications at PubMed
Contact Us
Contact Dr. Williams.

Deborah Winter
Associate Professor, Medicine (Rheumatology)
Computational immunology - using genomic approaches to study macrophages in disease

Deborah Winter
Research Area(s): Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics; Immunology
The goal of the Winter Lab of Macrophage Genomics is to apply genomic approaches to the role of immune cells in health and disease, particularly macrophages. Led by Dr. Deborah Winter, a computational immunologist, we employ the latest technologies for transcriptional and epigenomic assays, such as RNA-seq, ChIP-seq, ATAC-seq, single-cell, and spatial technologies, in combination with innovative quantitative approaches to analyze and model gene regulatory networks. Our goal is to understand how macrophages are programmed to maintain tissue homeostasis while priming for response to illness and injury. We are particularly interested in the role of macrophages in aging and autoimmune diseases such as scleroderma, rheumatoid arthritis and lupus. Our projects ranges from basic science with mouse models to translational research with clinical samples. A major goal of the lab is the application of functional genomic to a precision medicine approaches that can inform clinical decision making. We are committed to high standards for rigorous analysis and are continually adapting and optimizing computational techniques to address new challenges as they arise. We are currently recruiting highly motivated individuals with a variety of backgrounds to join the lab.
For more information, visit the faculty profile of Dr. Winter and Winter Lab site.
Publications
View Dr. Winter's publications at PubMed
Contact Us
Contact Dr. Winter at 312-503-0535

Gayle E Woloschak
Professor, Radiation Oncology, Radiology
Radiation-induced mutations in radiation-induced cancers; DNA-TiO2 nanoparticles; Radiosensitivity/motor neuron disease

Gayle E Woloschak
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Genetics, Epigenetics and Genomics
The Woloschak Lab members focus their research on three main areas.
The Janus Project: Studying radiation-induced mutations in radiation-induced cancers
This 30 year, $200 million set of experiments were performed at 150 laboratories and then terminated before the data were completely analyzed. Funded by the Department of Energy and National Aeronautic and Space Administration, department radiobiologists will continue the data analyses.
Members of the Woloschak laboratory have assumed responsibility from Argonne National Laboratory for archiving tissue associated with 30,000 mice and 4,000 dogs that received various doses and dose-rates of radiation.
These studies examined the effects of dose-rate on radiation-induced toxicities and radiation-induced cancer. They are analyzing cancer cells from these tissues to find differences in mutational spectra that occur in tumors induced in radiation-exposed animals compared to those that occur in spontaneous tumors. Recent scientific concerns about very low dose exposures makes this effort particularly important.
Collaborators
- University of Chicago
- Bundewehr Radiobiology Institute in Munich
- Argonne National Lab
DNA-TiO2 nanoparticles
The researchers have combined the functional properties of the biomolecule DNA and the inorganic compound TiO2. The project is oriented to investigating the functional use of these nanocomposites for intracellular manipulation, imaging and gene silencing.
Radiosensitivity/motor neuron disease
The project's purpose is to better understand the molecular basis for the combined abnormalities from a molecular-cellular perspective. Chip-based mRNA studies, gene promoter analyses, immunohistochemistry and standard molecular approaches are being used.
Learn More
For lab information and more, see Dr. Woloschak's faculty profile and Woloschak Lab site.
Publications
See Dr. Woloschak's publications on PubMed
Contact
Contact Dr. Woloschak at 312-503-4323

Waihay J Wong
Assistant Professor, Pathology
Clonal hematopoiesis, inflammation, and development of hematologic malignancy

Waihay J Wong
Research Area(s): Cancer Biology; Cell Biology; Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Immunology
My laboratory is interested in how gene mutations affect hematopoietic differentiation and downstream immune function. Specifically, we are interested in somatic mutations in common leukemia-causing genes occurring in the healthy aging population, a condition known as clonal hematopoiesis of indeterminate potential (CHIP). We employ state-of-the-art technologies, such as single-cell sequencing, high-resolution chromatin mapping, CRISPR screening, and transgenic mouse models to study the genetic and environmental contributions to hematopoietic myeloid cell expansion and convergence on inflammation.
A major focus of my research is to uncover the mechanistic association between hematopoietic development and innate inflammatory signaling. By dissecting the specific inflammatory responses in mutant myeloid-derived cells, we seek to gain a comprehensive understanding of the cellular and molecular mechanisms underlying myeloid expansion and progression to malignancy. Another focus is to further understand the significant risk association between CHIP and conditions of inflammaging based on identifying site-specific, spatially-dependent, immunological interactions between parenchymal cells and mutant myeloid clones. Through this work, we aim to identify therapeutic strategies to combat age-associated inflammation and myeloid neoplasia.
For more information, visit Dr. Wong's faculty profile.
Publications
View Dr. Wong's full list of publications in PubMed.
Contact

Yvette C Wong
Assistant Professor, Neurology (Movement Disorders)
Super-resolution live microscopy: Identifying new organelle dynamics underlying neurodegenerative diseases

Yvette C Wong
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Genetics, Epigenetics and Genomics; Neurobiology
Neurodegenerative diseases have been linked to the misregulation of multiple organelles including mitochondria and lysosomes, which are key for cellular and neuronal function. Moreover, organelles are highly dynamic and investigating their regulation in real-time is crucial for advancing our understanding of cell biology, cellular neuroscience, and neurodegenerative disease mechanisms. We recently identified mitochondria-lysosome contact sites as important regulators for mitochondrial and lysosomal network dynamics, which are implicated in multiple neurological diseases including Parkinson's and Charcot-Marie-Tooth disease.
Our ongoing research seeks to:
- Use super-resolution live cell microscopy to identify new cellular pathways
- Investigate how mitochondria-lysosome contact sites drive cellular & neuronal homeostasis and human disease pathogenesis
- Explore the roles and regulation of inter-organelle membrane contact sites
- Elucidate how organelle dynamics contribute to neurodegeneration in Parkinson’s, Charcot-Marie-Tooth, ALS & Alzheimer’s disease
For more information, visit the Yvette Wong Lab website or Yvette Wong's faculty profile.
Publications
View Dr. Wong's full list of publications in PubMed.
Contact Us

Jennifer D Wu
Professor, Urology, Microbiology-Immunology
Mechanisms of cancer immune evasion and development of novel immune therapy

Jennifer D Wu
Research Area(s): Cancer Biology; Immunology
Dr. Wu's goal to better understand prostate cancer and cancer in general and to find a better treatment for the untreatable terminal diseases, her current research focuses on the following aspects:
- Understanding the universal mechanisms of cancer immune evasion and development of novel cancer immunotherapy with specific focus on the NKG2D signaling pathways;
- Understanding why immunotherapy is not effective in prostate cancer and how to make it work with understanding underlying mechanisms of resistance;
- Discovering biomarkers to distinguish progressive vs. indolent prostate cancer to direct clinical decision making on treatment options;
- Understanding mechanism underlying immunotherapy (specifically, immune checkpoint inhibitor-therapy) induced toxicity and developing mechanism-direct personalized treatment for cancer patients to alleviate toxicity and to improve clinical outcomes.
For more information please view the faculty profile of Jennifer Wu, PhD.
Publications
View a full list of publications by Jennifer Wu at PubMed
Contact
Email Jennifer Wu, or phone at 312-503-1521

Rendong Yang
Associate Professor, Urology
Integrating genomics, big data, and computing to drive innovations in human genetics, precision oncology, and cancer immunotherapy

Rendong Yang
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics
The Yang laboratory is interested in the integrative analysis of large-scale, multi-dimensional genomic data to understand the initiation and progression of diseases. The research projects involve in the development of highly accurate and sensitive computational methods for analyzing large-scale genomic data, especially in the area of detecting and analyzing genetic variations and somatic mutations using next generation sequencing data. Current work in the lab is to explore the functional consequences of somatic alterations in cancer patients, to identify driver alterations, and to understand the genetic mechanisms of cancer progression and drug resistance by integrating multi-dimensional data from large-scale cancer studies such as The Cancer Genome Atlas (TCGA). Example projects span from technique-driven research that aims developing algorithms for a wide range of applications to hypothesis-driven investigation of specific biological problems where the main goal is the discovery and advancement of biological knowledge.
For more information visit Dr. Yang's lab website: https://ylab-hi.github.io/
Publications
See Dr. Yang's publications on PubMed.
Contact
Contact Dr. Yang.

Mee-Ngan F Yap
Associate Professor, Microbiology-Immunology
Molecular mechanisms of ribosome hibernation and antibiotic resistance

Mee-Ngan F Yap
Research Area(s): Cell Biology; Microbiology
We broadly investigate the mechanisms used in bacterial cells to regulate antibiotic resistance and gene expression at the translational level. Our long-term goals are to address mechanistic questions about ribosome specialization and resistance evolution, in addition to developing aptamer-based diagnostic tools for bacterial pathogens.
Current Projects
Multidrug resistant ribosome
Macrolides, lincosamides and streptogramins (MLS) are structurally distinct and broad-spectrum antibiotics that inhibit protein biosynthesis by binding to the 50S large subunit of bacterial ribosome. The efficacy of MLS has rapidly eroded due to the widespread dissemination of the Erm RNA methyltransferases that catalyze the transfer of two methyl groups to a conserved adenine nucleotide (m26A2058) in the 23S rRNA of the 50S subunit. This dimethylation sterically hinders the binding of all MLS antibiotics that share the overlapping A2058, in addition to abrogating the MLS resistant bacteria from host immune recognition. Our studies seek to addresses several unresolved questions: How is the expression of erm regulated under antibiotic selection? How does Erm find its target? What are the consequences of ribosome methylation? How do the next-generation antibiotics recognize the methylated ribosome?
Hibernating ribosome
The bacterial 100S ribosome (dimer of 70S complexes) is important for pathogenesis, translational repression, starvation responses, and ribosome turnover. Our goal is to establish a mechanistic understanding of the biogenesis and function of the 100S ribosome in translational silencing and staphylococcal pathogenesis. This project focuses on the following unexplored questions: What factors control the constitutive production of the 100S ribosome in S. aureus? Why are only specific mRNAs translationally repressed during ribosome hibernation? How is hibernation beneficial to ribosome stability? How is ribosome turnover linked to successful host colonization? These questions will be addressed through a multi-disciplinary approach that spans genetics, molecular biophysics, biochemistry and whole animal infection studies.
For lab information and more, see Dr. Yap's faculty profile and lab website.
Publications
See Dr. Yap's publications on ORCID.
Contact
Contact Dr. Yap.

Rui Yi
Professor, Pathology (Experimental Pathology), Dermatology
Mechanisms of skin development, stem cells, aging and cancer at the single-cell level

Rui Yi
Research Area(s): Cancer Biology; Cell Biology; Computational Biology and Bioinformatics; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics
Mammalian skin and its appendages function as the outermost barrier of the body to protect inner organs from environmental hazards and keep essential fluid within. Our research program studies mechanisms that govern cell fate specification, stem cell maintenance and aging as well as initiation and progression of cancer. We use single-cell genomics and computational tools, live animal imaging and genetically engineered mouse models to study gene expression regulation mediated by transcription factors, epigenetic regulators and post-transcriptional mechanisms mediated by miRNAs and RNA binding proteins at the single-cell resolution in mammalian skin.
Our research aims to address several fundamental questions in stem cell biology: how the developmental potential of embryonic progenitors and adult tissue stem cells is transmitted or restricted in their progenies at the molecular level when they go through critical transitions such as cell fate specification, self-renewal of stem cells as well as stress response, and how these regulatory mechanisms go awry in aging and diseases. Answers to these questions will help to manipulate skin stem cells for regenerative medicine and discover new treatment for human skin diseases.
View all lab publications via PubMed.
For more information, visit the faculty profile page of Rui Yi, PhD or visit the Yi Laboratory website.
Contact Us

Feng Yue
Professor, Biochemistry and Molecular Genetics, Pathology
Identify biomarkers and pathological variants in human cancers at single cell resolution

Feng Yue
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics
The long-term goal of Dr. Yue’s group is to use a combination of high throughput genomics, computational modeling, and functional assays to study how genetic variants contribute to the pathogenesis of human cancer. In particular, he is interested to identify the mutations that can disrupt the function of non-coding regulatory elements such as enhancers and further contribute to the pathogenesis of cancer. He has been actively involved with several large NIH-funded consortia, including the ENCODE Project, the 4D Nucleome consortium, and Impact of Genomic Variation on Function Consortium (IGVF).
For more information, please see Dr. Yue's faculty profile or the Yue lab website.
Publications
See Dr. Yue's publications on PubMed.
Contact
Contact Dr. Yue at 312-503-8248.

Bin Zhang
Professor, Medicine (Hematology and Oncology), Microbiology-Immunology, Pathology
Tumor-induced immune suppression

Bin Zhang
Research Area(s): Cancer Biology; Immunology; Pharmacology
Dr. Bin Zhang’s laboratory focuses on understanding how tumors evade immune surveillance and on developing strategies to restore effective anti-tumor immunity. His research centers on tumor-induced immune suppression, particularly the mechanisms by which cancer cells reprogram immune cells and the tumor microenvironment to inhibit T cells and foster tumor-promoting myeloid cell (e.g. neutrophil and macrophage) function based on the cutting-edge single-cell technical platforms and nanoengineering approaches.
The lab has made important contributions to identifying key immunosuppressive pathways—including metabolic and stress-response signaling—that allow tumors to escape immune destruction. Building on these mechanistic insights, Dr. Zhang’s group aims to translate their findings into novel immunotherapeutic strategies. Their work includes developing approaches to enhance the efficacy of immune checkpoint blockade, adoptive cell therapies, and combinatorial treatments that target both tumor cells and the immunosuppressive microenvironment. Ultimately, the lab seeks to improve cancer treatment outcomes by designing therapies that reinvigorate durable and robust anti-tumor immune responses.
For more information, visit the faculty profile page of Bin Zhang, MD/PhD.
Publications
View lab publications via Faculty Page.
Contact Us
Contact Dr. Zhang at 312-695-6180 or the Zhang Lab at 312-503-2435.

Ming Zhang
Associate Professor, Biochemistry and Molecular Genetics
Molecular Mechanisms of Tumorigenesis and Cancer Metastasis

Ming Zhang
Research Area(s): Cancer Biology; Cell Biology; Developmental and Stem Cell Biology; Immunology
The Zhang laboratory is focused on two research directions: 1) determining role of tumor suppressors in development and cancer progression and 2) identifying immune components that control breast cancer metastasis.
The main focus of my research program is to study the roles of tumor suppressors in normal development and in breast and prostate cancer progression, focusing on maspin and an Ets transcription factor PDEF. Maspin is a unique member of the SERPIN family that plays roles in normal tissue development, tumor metastasis and angiogenesis. Genetic studies by my laboratory using maspin transgenic and knockout mice demonstrated an important role of maspin in normal mammary, prostate and embryonic development. Recently, we have identified several new properties of maspin. As a protein that is present on cell surface, maspin controls cell-ECM adhesion. This function is responsible for maspin-mediated suppression of tumor cell motility and invasion. We have also discovered that maspin is involved in the induction of tumor cell apoptosis through a mitochondrial death pathway. The long-term goals of these projects are to elucidate the molecular mechanisms by which maspin and PDEF control tumor metastasis and to identify their physiological functions in development. These analyses are not only important for basic biology and but also may lead to a therapy for cancer and other developmental diseases.
Another focus of research in Zhang lab is to identify immune components that control breast cancer metastasis. Chronic inflammation not only increases neoplastic transformation but also drives the inhibition of the immune response in a protective negative-feedback mechanism. Suppressive immune cells are recruited to the sites of inflammation and function to inhibit both innate and adaptive immune responses, enabling tumor tolerance and unmitigated tumor progression. To study the interplay between tumor and immune cells, the Zhang lab has developed a unique animal model of breast cancer that reproduces different stages of breast cancer bone metastasis. Molecules that control tumor-immune cell interaction and immunosuppression have been identified. We are currently studying roles of these genes in tumor-driven evolution that control chronic inflammation and immunosuppression. We hypothesize that these key pro-inflammatory genes are upregulated during cancer progression, which function synergistically to recruit and activate suppressive MDSCs, TAMs and Tregs, inducing chronic inflammation and an immunosuppressive tumor microenvironment conducive to metastatic progression.
For more information visit Ming Zhang's faculty profile.
Publications
View publications by Ming Zhang in PubMed
Contact
Phone 312-503-0449

Peng Zhang
Assistant Professor, Neurological Surgery
Nano-hijacking tumor immunity for immunotherapy

Peng Zhang
Research Area(s): Cancer Biology; Immunology; Neurobiology; Pharmacology
Our lab has been focused on multidisciplinary research approaches, including nanotechnology, immunology, immune metabolism, cancer biology, and bioinformatics, to unveil the mechanisms of cancer therapy resistance, identify novel therapeutic targets, create innovative therapeutic strategies, and enable a bench-to-bedside translation.
One major research interest of our group is to create innovative therapeutic strategies for targeting immunosuppressive mechanisms in tumors. The immunosuppressive microenvironment is one major cause of cancer therapy resistance to both standard-of-care and immunotherapy. Tumor-associated myeloid cells (TAMCs) represent a predominant immune population and are a key driver of immunosuppression in different types of cancers. Building on our expertise in cancer immunology and nanotechnology, we have engineered innovative nano-therapeutics that enable effective in vivo targeting and modulation of TAMCs. Harnessing the functionality of TAMCs by impairing their immunosuppressive activity and/or remodeling them to be tumor-eradicating cells greatly boosts anti-tumor immunity and generates an immunologically “hot” tumor. We also seek to gain insight into the crosstalk between TAMCs and adaptive immunity (i.e. effector T & B cells).
Our lab has developed an antibody-directed precision targeting nanotechnology platform for precision in vivo targeting of immune cell subset of interest, such as macrophages, neutrophiles, dendritic cells, T cells, and B cells. It is a versatile system to deliver different types of therapeutics, including small molecule drugs and RNA therapeutics (e.g. mRNA & siRNA), enabling precision modulation of tumor-infiltrating immune cells. Our platforms serve as (i) new therapeutic & vaccination approaches to potentiate anti-tumor immunity and/or overcome therapy resistance in GBM, and (ii) powerful tools to advance mechanistic studies in basic immunology & biology for identifying new therapeutic targets.
Another key research interest of our laboratory is to dissect the interplay between standard-of-care and tumor immune microenvironment. i.e., how standard-of-care shapes tumor immune microenvironment, and how host immune response impacts the outcome of standard-of-care.
For more information, visit the Zhang lab website or Dr. Zhang's Faculty Profile page.
Publications
See Dr. Zhang's publications on PubMed
Contact
Email Dr. Zhang

Wei Zhang
Professor, Preventive Medicine
Genetics and epigenetics of complex traits including risks for common diseases and drug response

Wei Zhang
Research Area(s): Cancer Biology; Computational Biology and Bioinformatics; Genetics, Epigenetics and Genomics
Dr. Zhang is particularly interested in using high throughput technologies (e.g., microarray, next generation sequencing) and systems biology approaches to study the genetics of complex traits or phenotypes such as the risks of common diseases (e.g., cancer and lung disease), individual drug response and gene expression. Dr. Zhang is also interested in building bioinformatic databases that aim to provide user friendly access to primary data from pharmacogenomic and genome-wide association studies (GWAS). An on-going research interest of Dr. Zhang’s is the mapping of expression quantitative trait loci (eQTLs) in sarcoidosis and sickle cell disease, as well as the impact of eQTL mapping on the prioritization of GWAS results form these complex diseases.
For more information, visit Dr. Zhang's Faculty Profile page.
Publications
See Dr. Zhang's publications on PubMed
Contact
Email Dr. Zhang
Phone 312-503-1040

Ming Zhao
Associate Professor, Medicine (Cardiology)
Diagnostic markers and pathogenic mechanisms of human diseases based on changes in cellular membranes

Ming Zhao
Research Area(s): Biochemistry and Structural Biology; Cell Biology
Major research areas in the Zhao lab include:
- Apoptosis imaging technology development. Programmed cell death (apoptosis) plays a significant role in degenerative diseases. There is currently no clinical tool for assessing apoptosis in pathological conditions. Our research focuses on the development of optimal agents that combine sophisticated binding activities and favorable clearance kinetics for clinical translation.
- Assessing systemic toxicity in anticancer therapies. The outcome of chemotherapies hinges on the balance between tumor toxicity and patient tolerance. With the ability to noninvasively detect tissue apoptosis, we propose to assess anticancer therapies in a whole-body approach by monitoring tumor cell killing simultaneously with systemic tissue injury in response to chemotherapeutic agents. This is a transformative approach in oncology in terms of optimizing therapies on an individualized basis.
- Detecting myocardial injury in ischemic heart disease. Non-infarct myocardial injury in ischemic heart disease is of particular interest because this type of cardiac injury is not well understood in terms of its pathophysiological characteristics and its roles in long-term adverse cardiac events. Our research in this area focuses on the diagnosis of non-infarct myocardial injury, which in turn, will help address a significant gap in identifying patients at risk.
- Investigating the pathogenesis of antiphospholipid syndromes. The presence of circulating antibodies against phosphatidylethanolamine (PE) is positively correlated with clinical manifestations of antiphospholipid syndromes. However, the underlying pathogenic mechanism of anti-PE autoimmunity remains unknown. We have a major interest in investigating the cellular susceptibility to PE-binding agents, which in turn, will shed light on the potential pathogenic mechanism of aPE.
Publications
View publications by Dr. Zhao in PubMed.
Contact
Contact Dr. Zhao at 312-503-3226.

YouYang Zhao
Professor, Pediatrics (Critical Care), Pharmacology, Medicine (Pulmonary and Critical Care)
Molecular mechanisms of endothelial regeneration,resolution of inflammatory injury and endothelial and smooth muscle cell interaction in the pulmonary vascular diseases

YouYang Zhao
Research Area(s): Biochemistry and Structural Biology; Cancer Biology; Cardiovascular and Renal Biology; Cell Biology; Developmental and Stem Cell Biology; Genetics, Epigenetics and Genomics; Immunology; Lung Biology; Metabolism and Endocrinology; Pharmacology
Recovery of endothelial barrier integrity after vascular injury is vital for endothelial homeostasis and resolution of inflammation. Endothelial dysfunction plays a critical role in the initiation and progression of vascular diseases such as acute lung injury (ALI)/acute respiratory distress syndrome (ARDS) and atherosclerosis. A part of the research in the lab, employing genetically modified mouse models of human diseases, endothelial progenitor cells/stem cells, and translational research approach as well as nanomedicine, is to elucidate the molecular mechanisms of endothelial regeneration and resolution of inflammatory injury and determine how aging and epigenetics regulate these processes (J. Clin. Invest. 2006, 116: 2333; J. Exp. Med. 2010, 207:1675; Circulation 2016, 133: 2447). We are also studying the role of endothelial cells in regulating macrophage functional polarization and resolving inflammatory lung injury. These studies will identify druggable targets leading to novel therapeutic strategies to activate the intrinsic endothelial regeneration program to restore endothelial barrier integrity and reverse edema formation for the prevention and treatment of ARDS in patients.
Pulmonary hypertension is a progressive disease with poor prognosis and high mortality. We are currently investigating the molecular basis underlying the pathogenesis. We have recently identified the first mouse model of pulmonary arterial hypertension (PAH) with obliterative vascular remodeling including vascular occlusion and formation of plexiform-like lesions resembling the pathology of clinical PAH (Circulation 2016, 133: 2447). Our previous studies also show the critical role of oxidative/nitrative stress in the pathogenesis of PAH as seen in patients (PNAS 2002, 99:11375; J. Clin. Invest. 2009, 119: 2009). With these unique models and lung tissue and cells from idiopathic PAH patients, we will define the molecular and cellular mechanisms underlying severe vascular remodeling and provide novel therapeutic approaches for this devastating disease.
The Zhao lab employs the state-of-the art technologies including genetic lineage tracing, genetic depletion, genetic reporter, and CRISPR-mediated in vivo genomic editing as well as patient samples to study the molecular mechanisms of acute lung injury/ARDS, and pulmonary hypertension and identify novel therapeutics for these devastating diseases. Current studies include 1) molecular mechanisms of endothelial regeneration and vascular repair following inflammatory lung injury induced by sepsis and pneumonia; 2) how aging and epigenetics regulate this process; 3) how endothelial cells regulate macrophage and neuptrophil function for resolution of inflammation and host defense; 4) stem/progenitor cells in acute lung injury and pulmonary hypertension and cell-based therapy; 5) mechanisms of obliterative pulmonary vascular remodeling; 6) molecular basis of right heart failure; 7) pathogenic role of oxidative/nitrative stress; 8) lung regeneration; 9) drug discovery; 10) nanomedicine.
For more information, visit Dr. Zhao's Faculty Profile page
Publications
View publications by Youyang Zhao in PubMed.
Contact
Email Dr. Zhao

Jing Zheng
Professor, Otolaryngology - Head and Neck Surgery
Hearing and cochlear amplification, deafness-related proteins and cell death

Jing Zheng
Research Area(s): Biochemistry and Structural Biology; Cell Biology; Neurobiology
The goal of my lab is to identify and investigate molecules that play important roles in mammalian hearing, thus enriching our understanding of cochlear physiology and further developing a better strategy to prevent hearing loss. Deafness is commonly caused by defects in inner ear hair cells. In mammalian cochleae, inner hair cells (IHCs) function as sensory receptors conveying sound-related information to the central nervous system. Outer hair cells (OHCs) amplify the mechanical signals delivered to IHCs. The cooperation between IHCs and OHCs results in sensitive hearing and sharp tuning. Complex and sophisticated protein networks in hair cells facilitate their functions. Very often, genetic defects in a single protein can interfere with the entire network and cause deafness. Our research has been centered on several important proteins expressed in cochlea.
1. Molecular basis of cochlear amplification. OHCs undergo rapid somatic length changes when the voltage across their membrane is altered. This unique somatic electromotility provides the local mechanical amplification of the cochlear response to sound. Without OHCs, hearing threshold is elevated by 40-50 dB and frequency resolution deteriorates. Prestin is the motor protein of OHCs and is required for cochlear amplification (Zheng et al., Nature, 2000). Coincidently, prestin is only expressed in OHCs, which are also the most vulnerable cells in the organ of Corti. In the past, studying OHC amplification mechanisms and preventing OHC loss were considered two separate research fields. However, our recent data indicate a close connection between prestin's function and the vulnerability of OHCs to a variety of ototoxic exposures. To understand this link, we focus on investigating the molecular mechanism of the motor protein prestin using various cellular, biochemical and molecular biological methods.
2. Protein network of hair cells. We are focus on several deafness-related proteins: CDH23, CEACAM16 and Marshalin. Cadherin 23 is a tip-link protein of hair cells. CEACAM16 is an adhesive protein localized at the tectorial membrane (Zheng et al. PNAS, 2011). Marshalin is another newly identified microtubule minus-end binding protein that is expressed in the inner ear. Its expression is developmentally controlled (Zheng et al., Biology Open 2013). Very often, genetic defects in a single protein can interfere with the entire network and cause deafness. We are in the process of investigating interactions among these proteins and their physiological roles for normal hearing and deafness.
For more information, visit the faculty profile of Jing Zheng, PhD.
Publications
View Dr. Zheng's publications in PubMed
Contact
Phone 312-503-3417

Joshua J Ziarek
Associate Professor, Pharmacology
Molecular mechanisms of GPCR allostery for rational drug discovery

Joshua J Ziarek
Research Area(s): Biochemistry and Structural Biology; Computational Biology and Bioinformatics; Pharmacology
G protein-coupled receptors (GPCRs) dominate the therapeutic market – being the targets for more than 30 percent of FDA-approved drugs. They also recognize most drugs of abuse including opioids, cannabinoids, LSD, cocaine, and methamphetamine. Unlike many signaling proteins that function as binary switches between 'on and off' states, GPCR activity can be increased or decreased upon ligand binding - more akin to a dimmer switch. A single receptor may specifically recognize several ligands and respond uniquely to each, creating a complex conformational landscape. There is immense therapeutic potential in the ability to tune receptor signaling – and it remains poorly understood and virtually untapped.
The long-term vision of the Ziarek Lab is to decipher the molecular mechanisms of GPCR allostery for rational drug discovery. The field's current understanding of biased agonism derives primarily from cell-based functional assays that are easily complicated by receptor expression levels, off-target effects, cell type, and more. Integrating biophysical studies of purified protein with this cellular information would permit a complete molecular understanding. Some research areas the Ziarek lab is currently exploring are:
- designing novel GPCR ligands to combat cocaine, methamphetamine, and opioid substance abuse;
- investigating the conformational space of wildtype GPCRs using a combined biophysical and computational approach;
- characterizing the allosteric mechanisms of arrestin signaling;
- developing state-of-the-art nuclear magnetic resonance (NMR) probes, pulse programs and data analysis pipelines; and
- actualizing the full potential of NMR to explore GPCR conformational dynamics using receptors compatible with E. coli expression systems.
For more information, visit the faculty profile page of Joshua Ziarek or the Ziarek lab website.
Publications
View lab publications via PubMed.
Contact Us
Contact Dr. Ziarek
