Awarded Projects
The Polsky Urologic Cancer Institute of the Robert H. Lurie Comprehensive Cancer Center of Northwestern University at Northwestern Memorial Hospital is committed to driving transformative new insights and successes in urologic cancer research, education and discovery. To achieve this mission, the institute provides support for projects with promising potential to advance knowledge and foster new, innovative research in the field of urologic oncology.
We have selected five key projects to receive funding for the 2022 award year:
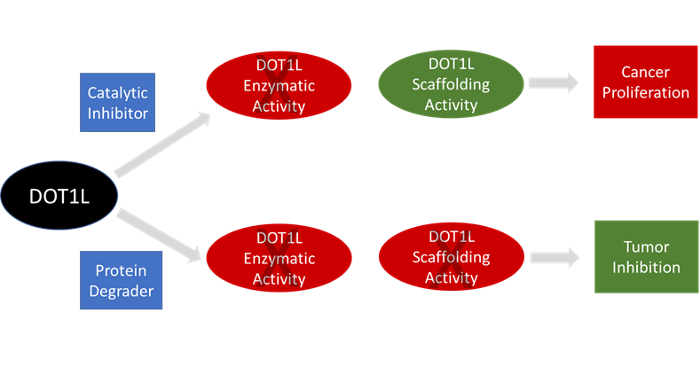
Targeted Degradation of DOT1L for Prostate Cancer Therapy
PI: Sarki Abdulkadir, MD, PhD, Professor of Urology, Feinberg School of Medicine
PI: Gary Schiltz, PhD, Research Professor, Center for Molecular Innovation and Drug Discovery, Northwestern University Chemistry of Life Processes Institute
The Abdulkadir lab recently identified the histone methyltransferase DOT1L as a major, targetable regulator of AR and MYC protein stability in therapy-resistant PC. However, therapeutic targeting of DOT1L with small-molecule inhibitors has thus far been unsuccessful. A major reason for this is because DOT1L, like other methyltransferases, possesses methyltransferase-independent functions. Thus, methyltransferase enzyme inhibitors may not fully suppress DOT1L function in cancer. To overcome these shortcomings, the lab developed a unique series of DOT1L PROTACS (proteolysis targeting chimeras) for targeted degradation of the DOT1L protein. In this project, the team will perform lead optimization of the novel DOT1L PROTACS for potency, selectivity, pharmacokinetics, pharmacodynamics, safety and anti-tumor efficacy properties. They expect to identify a clinical candidate DOT1L PROTAC that will enter IND-enabling studies.
Investigating Gender Differences in Mechanisms of Smoking-Related Carcinogenesis
PI: Joshua Meeks, MD, PhD, Edward M. Schaeffer, MD, PhD Professor of Urology; Assistant Professor of Urology and Biochemistry and Molecular Genetics, Feinberg School of Medicine
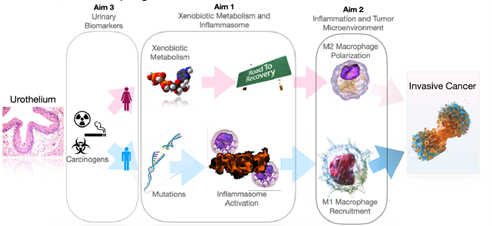
Men are 3-4-times more likely to develop bladder cancer than women. Yet, women have worse survival at every stage with up to a 50% increased risk of death from bladder cancer. Dr. Meeks is investigating gender differences in carcinogenesis to identify the causes of cancer initiation with the long-term goal to develop therapeutic strategies to prevent or screen patients at risk for bladder cancer.
Dr. Meeks discovered a strong link between hormone regulation of the inflammasome pathway in males that is believed to be responsible for amplifying carcinogenesis. When women go through menopause, they too become susceptible for increased inflammation after carcinogen exposure. In this project, is investigating how bladder cancer starts. In particular, he will further dissect this mechanism in cells and mice and evaluating urine as biomarker for hormonally regulated inflammatory responseDeveloping Digital Pathology and Artificial Intelligence for Prostate Cancer Risk Stratification
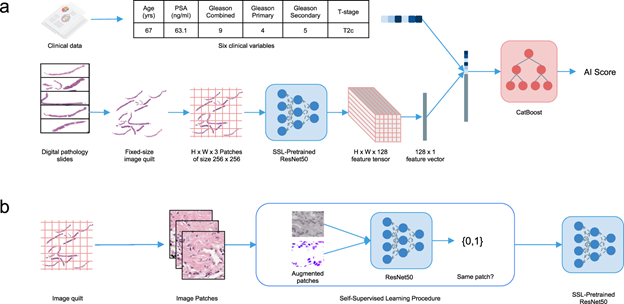
PI: Ashley Ross, MD, PhD, Associate Professor of Urology, Feinberg School of Medicine
Figure adapted from Esteva et al NPJ Digital Medicine 2022
Prostate cancer is the most common non-skin cancer in men and is the second most common cause of male cancer death. Disease risk varies widely with some cancers being slow growing and not needing treatment and others being aggressive and life threatening. Identification and characterization of disease under the microscope by pathologists has long been a foundational element for our clinical decision making to treat men with prostate cancer.
While powerful, human pathological tumor classification is limited by inter-observer variability. Emphasis is on cancer cells only with the environment around the cancer being largely ignored. Accuracy requires expert genitourinary pathologists, and the process of manual review is cumbersome, time consuming, and subjective.
Increases in computing power and data storage now allow for digitization of pathology and analysis by deep learning. Dr. Ross and his team aim to develop an inter-departmental resource of annotated digital pathology from men with prostate cancer. The team will utilize subsets of their digital library to further develop, validate and iterate quantitative artificial intelligence derived pathological grading and test newly developed deep learning based algorithms for their ability to predict disease outcomes. When successful, beyond forming a rich resource for future research, the advances made will be immediately translatable into clinical workflow where it could increase efficiency and improve risk stratification, and hence prostate cancer patient outcomes.
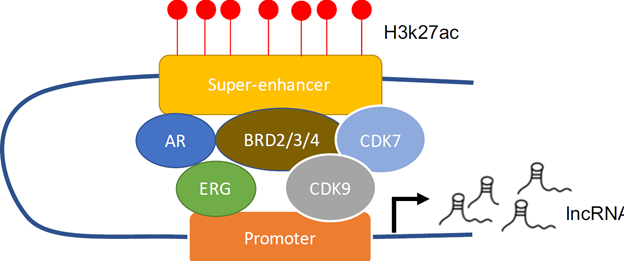
Delineating Superenhancer-Regulated lncRNA Drivers in Metastatic Prostate Cancer
PI: Rendong Yang, PhD, Associate Professor of Urology, Feinberg School of Medicine
Prostate cancer is most often found in early stages, which means the cancer cells remain in the prostate so the disease is highly treatable with surgery or radiation therapy. However, after localized therapy, some patients progress to an aggressive disease where the cancer has spread beyond the original area to other parts of the body. Although there are tests available to efficiently diagnose the early stage prostate cancer, these tests cannot predict whether the patients will progress to an advanced stage. On the other hand, patients with advanced stage prostate cancer will receive androgen deprivation therapy (ADT). However, most of the patients under ADT will eventually develop resistance to this therapy, which is called castration resistant prostate cancer (CRPC), leaving physicians with no options to counteract the inevitable. Therefore, there is an urgent need to (1) find novel biomarkers that can be used to identify patients that will progress to advanced disease; (2) identify novel targets that are druggable for CRPC patients.
Recent advances in high-throughput sequencing technologies have revealed that over 70% of the human genome is actively expressed. Whereas, only ~2% of the genome are coding to make proteins, the remaining is expressed as noncoding RNAs. A class of noncoding RNAs which are larger than 200bp called long noncoding RNA (lncRNAs) has been discovered to play important roles in cancer progression.
In this project, Dr. Yang will utilize cutting-edge technologies to test hypotheses about the role of a class of non-protein-coding RNA (lncRNA) in the development of metastatic CRPC. Identifying these novel lncRNA will help design better tools that can accurately predict the progression of cancer as well as novel therapeutic interventions that can target advanced tumors.
Targeting Metabolic Adaptation to Treat TP53-mutated Lethal Prostate Cancer
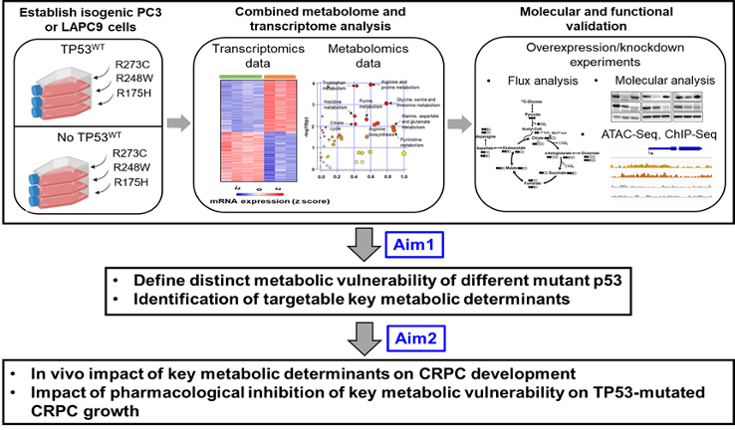
PI: Young Yoo, PhD, Research Assistant Professor of Urology, Feinberg School of Medicine
Alteration in the TP53 gene as gain-of-function (GOF) missense mutations is one of the most common events responsible for treatment resistance in prostate cancer. However, no drugs have been developed to date that directly target mutant TP53, so these lethal tumors remain untreatable.
Tumor cells require huge amounts of nutrients to keep up with their rapid malignant growth and mutant TP53 plays an important role in remodeling tumor metabolism to produce enough energy and nutrients to sustain their growth and survival. Therefore, oncogenic action of mutant TP53 could make tumors more susceptible to metabolic inhibition. However, all TP53 mutations are not equal and respond differently to treatments. In addition, TP53 mutations also often exist in the presence or absence of wild type TP53, which result in distinct oncogenic capability of cancer cells. Accordingly, it is of great importance to uncover the differential effects of individual mutant p53 proteins on metabolic vulnerabilities because it could contribute to developing more effective therapeutics targeting mutant TP53 and metabolic vulnerability to treat lethal prostate cancer patients with limited treatment options.
In this research, Dr. Yoo will dissect the metabolism of TP53 mutation variants through novel CRPC models mimicking human tumor mutation to identify potential vulnerabilities that can be targeted therapeutically and exploited to improve prostate cancer outcomes. Successful completion of these studies will provide a novel biological insight into the molecular link between TP53 mutation types and metabolic dependency, providing metabolic landscape associated with TP53 mutation in prostate cancer. Thus, targeting this metabolic pathway would be a new avenue to treat lethal prostate cancers with TP53 alteration.